A Network-Based Analysis Reveals the Mechanism Underlying Vitamin D in Suppressing Cytokine Storm and Virus in SARS-CoV-2 Infection
- PMID: 33362771
- PMCID: PMC7756074
- DOI: 10.3389/fimmu.2020.590459
A Network-Based Analysis Reveals the Mechanism Underlying Vitamin D in Suppressing Cytokine Storm and Virus in SARS-CoV-2 Infection
Abstract
Background: SARS-CoV-2 causes ongoing pandemic coronavirus disease of 2019 (COVID-19), infects the cells of the lower respiratory tract that leads to a cytokine storm in a significant number of patients resulting in severe pneumonia, shortness of breathing, respiratory and organ failure. Extensive studies suggested the role of Vitamin D in suppressing cytokine storm in COVID-19 and reducing viral infection; however, the precise molecular mechanism is not clearly known. In this work, bioinformatics and systems biology approaches were used to understand SARS-CoV-2 induced cytokine pathways and the potential mechanism of Vitamin D in suppressing cytokine storm and enhancing antiviral response.
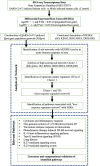
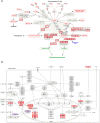
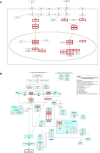
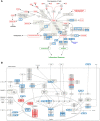
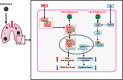
Results: This study used transcriptome data and identified 108 differentially expressed host genes (DEHGs) in SARS-CoV-2 infected normal human bronchial epithelial (NHBE) cells compared to control. Then, the DEHGs was integrated with the human protein-protein interaction data to generate a SARS-CoV-2 induced host gene regulatory network (SiHgrn). Analysis of SiHgrn identified a sub-network "Cluster 1" with the highest MCODE score, 31 up-regulated genes, and predominantly associated immune and inflammatory response. Interestingly, the iRegulone tool identified that "Cluster 1" is under the regulation of transcription factors STAT1, STAT2, STAT3, POU2F2, and NFkB1, collectively referred to as "host response signature network". Functional enrichment analysis with NDEx revealed that the "host response signature network" is predominantly associated with critical pathways, including "cytokines and inflammatory response", "non-genomic action of Vitamin D", "the human immune response to tuberculosis", and "lung fibrosis". Finally, in-depth analysis and literature mining revealed that Vitamin D binds with its receptor and could work through two different pathways: (i) it inhibits the expression of pro-inflammatory cytokines through blocking the TNF induced NFkB1 signaling pathway; and (ii) it initiates the expression of interferon-stimulating genes (ISGs) for antiviral defense program through activating the IFN-α induced Jak-STAT signaling pathway.
Conclusion: This comprehensive study identified the pathways associated with cytokine storm in SARS-CoV-2 infection. The proposed underlying mechanism of Vitamin D could be promising in suppressing the cytokine storm and inducing a robust antiviral response in severe COVID-19 patients. The finding in this study urgently needs further experimental validations for the suitability of Vitamin D in combination with IFN-α to control severe COVID-19.
Keywords: COVID-19; RNA sequencing; SARS-CoV-2; bioinformatics; cytokine storm; lung fibrosis; regulatory network; vitamin D.
Copyright © 2020 Ahmed.
Conflict of interest statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
A Computational Approach Identified Andrographolide as a Potential Drug for Suppressing COVID-19-Induced Cytokine Storm.Front Immunol. 2021 Jun 24;12:648250. doi: 10.3389/fimmu.2021.648250. eCollection 2021. Front Immunol. 2021. PMID: 34248936 Free PMC article.
-
Cytokine storm modulation in COVID-19: a proposed role for vitamin D and DPP-4 inhibitor combination therapy (VIDPP-4i).Immunotherapy. 2021 Jun;13(9):753-765. doi: 10.2217/imt-2020-0349. Epub 2021 Apr 28. Immunotherapy. 2021. PMID: 33906375 Free PMC article.
-
Putative roles of vitamin D in modulating immune response and immunopathology associated with COVID-19.Virus Res. 2021 Jan 15;292:198235. doi: 10.1016/j.virusres.2020.198235. Epub 2020 Nov 21. Virus Res. 2021. PMID: 33232783 Free PMC article. Review.
-
Janus kinase signaling as risk factor and therapeutic target for severe SARS-CoV-2 infection.Eur J Immunol. 2021 May;51(5):1071-1075. doi: 10.1002/eji.202149173. Epub 2021 Mar 22. Eur J Immunol. 2021. PMID: 33675065 Free PMC article. Review.
-
Dysregulated Interferon Response and Immune Hyperactivation in Severe COVID-19: Targeting STATs as a Novel Therapeutic Strategy.Front Immunol. 2022 May 17;13:888897. doi: 10.3389/fimmu.2022.888897. eCollection 2022. Front Immunol. 2022. PMID: 35663932 Free PMC article. Review.
Cited by
-
Response to Dr. DF Naude.Explore (NY). 2022 Mar-Apr;18(2):150-151. doi: 10.1016/j.explore.2021.10.002. Epub 2021 Oct 2. Explore (NY). 2022. PMID: 34642105 Free PMC article. No abstract available.
-
The role of peroxisome proliferator-activated receptors in the modulation of hyperinflammation induced by SARS-CoV-2 infection: A perspective for COVID-19 therapy.Front Immunol. 2023 Feb 17;14:1127358. doi: 10.3389/fimmu.2023.1127358. eCollection 2023. Front Immunol. 2023. PMID: 36875108 Free PMC article. Review.
-
Inflammatory reflex disruption in COVID-19.Clin Exp Neuroimmunol. 2022 Apr 29:10.1111/cen3.12703. doi: 10.1111/cen3.12703. Online ahead of print. Clin Exp Neuroimmunol. 2022. PMID: 35600135 Free PMC article. Review.
-
A Narrative Review on the Potential Role of Vitamin D3 in the Prevention, Protection, and Disease Mitigation of Acute and Long COVID-19.Curr Nutr Rep. 2023 Jun;12(2):215-223. doi: 10.1007/s13668-023-00471-2. Epub 2023 May 5. Curr Nutr Rep. 2023. PMID: 37145350 Free PMC article. Review.
-
Periodontal Disease and Other Adverse Health Outcomes Share Risk Factors, including Dietary Factors and Vitamin D Status.Nutrients. 2023 Jun 17;15(12):2787. doi: 10.3390/nu15122787. Nutrients. 2023. PMID: 37375691 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

