The complete chloroplast genome of Tainia dunnii (Orchidaceae): genome structure and evolution
- PMID: 33366394
- PMCID: PMC7720984
- DOI: 10.1080/23802359.2019.1693935
The complete chloroplast genome of Tainia dunnii (Orchidaceae): genome structure and evolution
Abstract
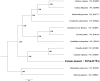
Tainia dunnii is a terrestrial orchid with high ornamental value. Herein, we assembled the complete chloroplast genome of Tainia dunnii by next-generation sequencing technologies. The complete chloroplast genome sequence of Tainia dunnii is 158,305 base pairs (bp) in length, including a pair of inverted repeat regions (IRs, 25,244 bp), one large single-copy region (LSC, 86,819 bp), one small single-copy region (SSC, 20,998 bp). Besides, the complete chloroplast genome contains 136 genes in total, including 88 protein-coding genes, 38 tRNA genes, and 8 rRNA genes. Phylogenetic analysis showed that Tainia dunnii has the closest relationship with Calanthe davidii and Calanthe triplicata. Our study lay a foundation for further research of Tainia dunnii.
Keywords: Chloroplast genome; Orchidaceae; Tainia dunnii; phylogenetic analysis.
© 2019 The Author(s). Published by Informa UK Limited, trading as Taylor & Francis Group.
Conflict of interest statement
No potential conflict of interest was reported by the authors.
Figures
References
-
- Jin JJ, Yu WB, Yang JB, Song Y, Yi TS, Li DZ. 2018. GetOrganelle: a simple and fast pipeline for de novo assembly of a complete circular chloroplast genome using genome skimming data. bioRxiv. 256479.
LinkOut - more resources
Full Text Sources