Chromosomal-Level Genome Assembly of Silver Sillago (Sillago sihama)
- PMID: 33367716
- PMCID: PMC7875006
- DOI: 10.1093/gbe/evaa272
Chromosomal-Level Genome Assembly of Silver Sillago (Sillago sihama)
Abstract
Silver sillago, Sillago sihama is a member of the family Sillaginidae and found in all Chinese inshore waters. It is an emerging commercial marine aquaculture species in China. In this study, high-quality chromosome-level reference genome of S. sihama was first constructed using PacBio Sequel sequencing and high-throughput chromosome conformation capture (Hi-C) technique. A total of 66.16 Gb clean reads were generated by PacBio sequencing platforms. The genome-scale was 521.63 Mb with 556 contigs, and 13.54 Mb of contig N50 length. Additionally, Hi-C scaffolding of the genome resulted in 24 chromosomes containing 96.93% of the total assembled sequences. A total of 23,959 protein-coding genes were predicted in the genome, and 96.51% of the genes were functionally annotated in public databases. A total of 71.86 Mb repetitive elements were detected, accounting for 13.78% of the genome. The phylogenetic relationships of silver sillago with other teleosts showed that silver sillago was separated from the common ancestor of Sillago sinica ∼7.92 Ma. Comparative genomic analysis of silver sillago with other teleosts showed that 45 unique and 100 expansion gene families were identified in silver sillago. In this study, the genomic resources provide valuable reference genomes for functional genomics research of silver sillago.
Keywords: Hi-C; PacBio; chromosomal assembly; genome; silver sillago.
© The Author(s) 2020. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures
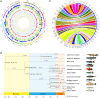
References
-
- Alioto T, Blanco E, Parra G, Guigó R. 2018. Using geneid to identify genes. Curr Protoc Bioinformatics. 64(1):e56. - PubMed
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. 1990. Basic local alignment search tool. J Mol Biol. 215(3):403–410. - PubMed
-
- Burge C, Karlin S. 1997. Prediction of complete gene structures in human genomic DNA. J Mol Biol. 268(1):78–94. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

