Evolution toward beta common chain receptor usage links the matrix proteins of HIV-1 and its ancestors to human erythropoietin
- PMID: 33372148
- PMCID: PMC7812818
- DOI: 10.1073/pnas.2021366118
Evolution toward beta common chain receptor usage links the matrix proteins of HIV-1 and its ancestors to human erythropoietin
Abstract
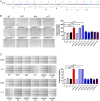
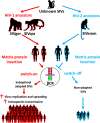
The HIV-1 matrix protein p17 (p17) is a pleiotropic molecule impacting on different cell types. Its interaction with many cellular proteins underlines the importance of the viral protein as a major determinant of human specific adaptation. We previously showed the proangiogenic capability of p17. Here, by integrating functional analysis and receptor binding, we identify a functional epitope that displays molecular mimicry with human erythropoietin (EPO) and promotes angiogenesis through common beta chain receptor (βCR) activation. The functional EPO-like epitope was found to be present in the matrix protein of HIV-1 ancestors SIV originated in chimpanzees (SIVcpz) and gorillas (SIVgor) but not in that of HIV-2 and its ancestor SIVsmm from sooty mangabeys. According to biological data, evolution of the EPO-like epitope showed a clear differentiation between HIV-1/SIVcpz-gor and HIV-2/SIVsmm branches, thus highlighting this epitope on p17 as a divergent signature discriminating HIV-1 and HIV-2 ancestors. P17 is known to enhance HIV-1 replication. Similarly to other βCR ligands, p17 is capable of attracting and activating HIV-1 target cells and promoting a proinflammatory microenvironment. Thus, it is tempting to speculate that acquisition of an epitope on the matrix proteins of HIV-1 ancestors capable of triggering βCR may have represented a critical step to enhance viral aggressiveness and early human-to-human SIVcpz/gor dissemination. The hypothesis that the p17/βCR interaction and βCR abnormal stimulation may also play a role in sustaining chronic activation and inflammation, thus marking the difference between HIV-1 and HIV-2 in term of pathogenicity, needs further investigation.
Keywords: HIV-1 and HIV-2 ancestors; HIV-1 evolutionary trajectory; HIV-1 matrix protein p17; common beta chain receptor; human erythropoietin.
Copyright © 2021 the Author(s). Published by PNAS.
Conflict of interest statement
Competing interest statement: F.C. and A.C. are listed as inventors in a patent application related to this study.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

