Chemoinformatics-based enumeration of chemical libraries: a tutorial
- PMID: 33372622
- PMCID: PMC7590480
- DOI: 10.1186/s13321-020-00466-z
Chemoinformatics-based enumeration of chemical libraries: a tutorial
Abstract
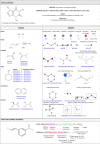
Virtual compound libraries are increasingly being used in computer-assisted drug discovery applications and have led to numerous successful cases. This paper aims to examine the fundamental concepts of library design and describe how to enumerate virtual libraries using open source tools. To exemplify the enumeration of chemical libraries, we emphasize the use of pre-validated or reported reactions and accessible chemical reagents. This tutorial shows a step-by-step procedure for anyone interested in designing and building chemical libraries with or without chemoinformatics experience. The aim is to explore various methodologies proposed by synthetic organic chemists and explore affordable chemical space using open-access chemoinformatics tools. As part of the tutorial, we discuss three examples of design: a Diversity-Oriented-Synthesis library based on lactams, a bis-heterocyclic combinatorial library, and a set of target-oriented molecules: isoindolinone based compounds as potential acetylcholinesterase inhibitors. This manuscript also seeks to contribute to the critical task of teaching and learning chemoinformatics.
Keywords: Chemical enumeration; Chemoinformatics; Combinatorial libraries; DOS synthesis; Drug design; Education; KNIME; Python.
Conflict of interest statement
The authors have declared no competing interest.
Figures





References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

