Active-Site Controlled, Jahn-Teller Enabled Regioselectivity in Reductive S-C Bond Cleavage of S-Adenosylmethionine in Radical SAM Enzymes
- PMID: 33372786
- PMCID: PMC7934139
- DOI: 10.1021/jacs.0c10925
Active-Site Controlled, Jahn-Teller Enabled Regioselectivity in Reductive S-C Bond Cleavage of S-Adenosylmethionine in Radical SAM Enzymes
Abstract
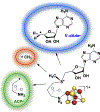
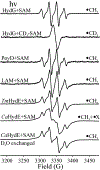
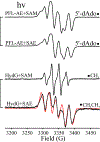
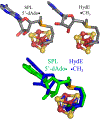
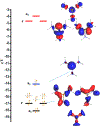
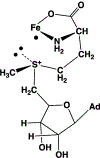
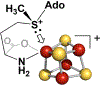
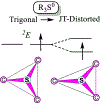
Catalysis by canonical radical S-adenosyl-l-methionine (SAM) enzymes involves electron transfer (ET) from [4Fe-4S]+ to SAM, generating an R3S0 radical that undergoes regioselective homolytic reductive cleavage of the S-C5' bond to generate the 5'-dAdo· radical. However, cryogenic photoinduced S-C bond cleavage has regioselectively yielded either 5'-dAdo· or ·CH3, and indeed, each of the three SAM S-C bonds can be regioselectively cleaved in an RS enzyme. This diversity highlights a longstanding central question: what controls regioselective homolytic S-C bond cleavage upon SAM reduction? We here provide an unexpected answer, founded on our observation that photoinduced S-C bond cleavage in multiple canonical RS enzymes reveals two enzyme classes: in one, photolysis forms 5'-dAdo·, and in another it forms ·CH3. The identity of the cleaved S-C bond correlates with SAM ribose conformation but not with positioning and orientation of the sulfonium center relative to the [4Fe-4S] cluster. We have recognized the reduced-SAM R3S0 radical is a (2E) state with its antibonding unpaired electron in an orbital doublet, which renders R3S0 Jahn-Teller (JT)-active and therefore subject to vibronically induced distortion. Active-site forces induce a JT distortion that localizes the odd electron in a single priority S-C antibond, which undergoes regioselective cleavage. In photolytic cleavage those forces act through control of the ribose conformation and are transmitted to the sulfur via the S-C5' bond, but during catalysis thermally induced conformational changes that enable ET from a cluster iron generate dominant additional forces that specifically select S-C5' for cleavage. This motion also can explain how 5'-dAdo· subsequently forms the organometallic intermediate Ω.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

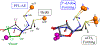
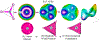

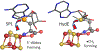
Similar articles
-
ENDOR Spectroscopy Reveals the "Free" 5'-Deoxyadenosyl Radical in a Radical SAM Enzyme Active Site Actually is Chaperoned by Close Interaction with the Methionine-Bound [4Fe-4S]2+ Cluster.J Am Chem Soc. 2024 Feb 14;146(6):3710-3720. doi: 10.1021/jacs.3c09428. Epub 2024 Feb 3. J Am Chem Soc. 2024. PMID: 38308759 Free PMC article.
-
Photoinduced Electron Transfer in a Radical SAM Enzyme Generates an S-Adenosylmethionine Derived Methyl Radical.J Am Chem Soc. 2019 Oct 9;141(40):16117-16124. doi: 10.1021/jacs.9b08541. Epub 2019 Sep 26. J Am Chem Soc. 2019. PMID: 31509404 Free PMC article.
-
Mechanism of Radical Initiation in the Radical SAM Enzyme Superfamily.Annu Rev Biochem. 2023 Jun 20;92:333-349. doi: 10.1146/annurev-biochem-052621-090638. Epub 2023 Apr 4. Annu Rev Biochem. 2023. PMID: 37018846 Free PMC article. Review.
-
Radical SAM Enzyme Spore Photoproduct Lyase: Properties of the Ω Organometallic Intermediate and Identification of Stable Protein Radicals Formed during Substrate-Free Turnover.J Am Chem Soc. 2020 Oct 28;142(43):18652-18660. doi: 10.1021/jacs.0c08585. Epub 2020 Oct 15. J Am Chem Soc. 2020. PMID: 32966073 Free PMC article.
-
Radical SAM enzymes: Nature's choice for radical reactions.FEBS Lett. 2023 Jan;597(1):92-101. doi: 10.1002/1873-3468.14519. Epub 2022 Oct 27. FEBS Lett. 2023. PMID: 36251330 Free PMC article. Review.
Cited by
-
Mechanism of Radical S-Adenosyl-l-methionine Adenosylation: Radical Intermediates and the Catalytic Competence of the 5'-Deoxyadenosyl Radical.J Am Chem Soc. 2022 Mar 23;144(11):5087-5098. doi: 10.1021/jacs.1c13706. Epub 2022 Mar 8. J Am Chem Soc. 2022. PMID: 35258967 Free PMC article.
-
Initiation, Propagation, and Termination in the Chemistry of Radical SAM Enzymes.Biochemistry. 2024 Dec 17;63(24):3161-3183. doi: 10.1021/acs.biochem.4c00518. Epub 2024 Dec 3. Biochemistry. 2024. PMID: 39626071 Free PMC article. Review.
-
The B12-independent glycerol dehydratase activating enzyme from Clostridium butyricum cleaves SAM to produce 5'-deoxyadenosine and not 5'-deoxy-5'-(methylthio)adenosine.J Inorg Biochem. 2022 Feb;227:111662. doi: 10.1016/j.jinorgbio.2021.111662. Epub 2021 Nov 12. J Inorg Biochem. 2022. PMID: 34847521 Free PMC article.
-
HydG, the "dangler" iron, and catalytic production of free CO and CN-: implications for [FeFe]-hydrogenase maturation.Dalton Trans. 2021 Aug 4;50(30):10405-10422. doi: 10.1039/d1dt01359a. Dalton Trans. 2021. PMID: 34240096 Free PMC article.
-
ENDOR Spectroscopy Reveals the "Free" 5'-Deoxyadenosyl Radical in a Radical SAM Enzyme Active Site Actually is Chaperoned by Close Interaction with the Methionine-Bound [4Fe-4S]2+ Cluster.J Am Chem Soc. 2024 Feb 14;146(6):3710-3720. doi: 10.1021/jacs.3c09428. Epub 2024 Feb 3. J Am Chem Soc. 2024. PMID: 38308759 Free PMC article.
References
-
- Frey PA; Hegeman AD; Ruzicka FJ The Radical SAM Superfamily. Crit. Rev. Biochem. Mol. Biol 2008, 43 (1), 63–88. - PubMed
-
- Nicolet Y Structure–function relationships of radical SAM enzymes. Nature Catalysis 2020, 3 (4), 337–350.
-
- Landgraf BJ; McCarthy EL; Booker SJ Radical S-Adenosylmethionine Enzymes in Human Health and Disease. Annu. Rev. Biochem 2016, 85 (1), 485–514. - PubMed
-
- Holliday GL; Akiva E; Meng EC; Brown SD; Calhoun S; Pieper U; Sali A; Booker SJ; Babbitt PC Chapter One—Atlas of the Radical SAM Superfamily: Divergent Evolution of Function Using a “Plug and Play” Domain. In Methods in Enzymology; Bandarian V, Ed.; Academic Press: 2018; Vol. 606, pp 1–71. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

