Modulation of Aub-TDRD interactions elucidates piRNA amplification and germplasm formation
- PMID: 33376130
- PMCID: PMC7772777
- DOI: 10.26508/lsa.202000912
Modulation of Aub-TDRD interactions elucidates piRNA amplification and germplasm formation
Abstract
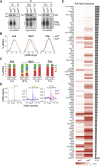
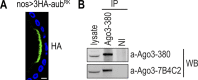
Aub guided by piRNAs ensures genome integrity by cleaving retrotransposons, and genome propagation by trapping mRNAs to form the germplasm that instructs germ cell formation. Arginines at the N-terminus of Aub (Aub-NTRs) interact with Tudor and other Tudor domain-containing proteins (TDRDs). Aub-TDRD interactions suppress active retrotransposons via piRNA amplification and form germplasm via generation of Aub-Tudor ribonucleoproteins. Here, we show that Aub-NTRs are dispensable for primary piRNA biogenesis but essential for piRNA amplification and that their symmetric dimethylation is required for germplasm formation and germ cell specification but largely redundant for piRNA amplification.
© 2020 Vrettos et al.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures






References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
