Design of a companion bioinformatic tool to detect the emergence and geographical distribution of SARS-CoV-2 Spike protein genetic variants
- PMID: 33380328
- PMCID: PMC7772798
- DOI: 10.1186/s12967-020-02675-4
Design of a companion bioinformatic tool to detect the emergence and geographical distribution of SARS-CoV-2 Spike protein genetic variants
Abstract
Background: Tracking the genetic variability of Severe Acute Respiratory Syndrome CoronaVirus 2 (SARS-CoV-2) is a crucial challenge. Mainly to identify target sequences in order to generate robust vaccines and neutralizing monoclonal antibodies, but also to track viral genetic temporal and geographic evolution and to mine for variants associated with reduced or increased disease severity. Several online tools and bioinformatic phylogenetic analyses have been released, but the main interest lies in the Spike protein, which is the pivotal element of current vaccine design, and in the Receptor Binding Domain, that accounts for most of the neutralizing the antibody activity.
Methods: Here, we present an open-source bioinformatic protocol, and a web portal focused on SARS-CoV-2 single mutations and minimal consensus sequence building as a companion vaccine design tool. Furthermore, we provide immunogenomic analyses to understand the impact of the most frequent RBD variations.
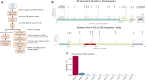
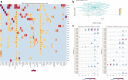
Results: Results on the whole GISAID sequence dataset at the time of the writing (October 2020) reveals an emerging mutation, S477N, located on the central part of the Spike protein Receptor Binding Domain, the Receptor Binding Motif. Immunogenomic analyses revealed some variation in mutated epitope MHC compatibility, T-cell recognition, and B-cell epitope probability for most frequent human HLAs.
Conclusions: This work provides a framework able to track down SARS-CoV-2 genomic variability.
Keywords: Bioinformatic workflow; COVID mutations; Docker; SARS-CoV-2 genome; SARS-CoV-2 mutation; SARS-CoV-2 vaccine.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

References
-
- Koyama T, Platt DE, Parida L. Variant analysis of COVID-19 genomes. J Bull World Heal Organ. 2020;2:1–21. https://www.researchgate.net/publication/339461351_Variant_analysis_of_C... - PMC - PubMed
-
- Chiara M, Horner DS, Pesole G. Comparative genomics suggests limited variability and similar evolutionary patterns between major clades of SARS-Cov-2. bioRxiv. 2020 doi: 10.1101/2020.03.30.016790. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

