HeartBioPortal2.0: new developments and updates for genetic ancestry and cardiometabolic quantitative traits in diverse human populations
- PMID: 33382884
- PMCID: PMC7774463
- DOI: 10.1093/database/baaa115
HeartBioPortal2.0: new developments and updates for genetic ancestry and cardiometabolic quantitative traits in diverse human populations
Abstract
Cardiovascular disease (CVD) is the leading cause of death worldwide for all genders and across most racial and ethnic groups. However, different races and ethnicities exhibit different rates of CVD and its related cardiorenal and metabolic comorbidities, suggesting differences in genetic predisposition and risk of onset, as well as socioeconomic and lifestyle factors (diet, exercise, etc.) that act upon an individual's unique underlying genetic background. Here, we present HeartBioPortal2.0, a major update to HeartBioPortal, the world's largest CVD genetics data precision medicine platform for harmonized CVD-relevant genetic variants, which now enables search and analysis of human genetic information related to heart disease across ethnically diverse populations and cardiovascular/renal/metabolic quantitative traits pertinent to CVD pathophysiology. HeartBioPortal2.0 is structured as a cloud-based computing platform and knowledge portal that consolidates a multitude of CVD-relevant genomic data modalities into a single powerful query and browsing interface between data and user via a user-friendly web application publicly available to the scientific research community. Since its initial release, HeartBioPortal2.0 has added new cardiovascular/renal/metabolic disease-relevant gene expression data as well as genetic association data from numerous large-scale genome-wide association study consortiums such as CARDIoGRAMplusC4D, TOPMed, FinnGen, AFGen, MESA, MEGASTROKE, UK Biobank, CHARGE, Biobank Japan and MyCode, among other studies. In addition, HeartBioPortal2.0 now includes support for quantitative traits and ethnically diverse populations, allowing users to investigate the shared genetic architecture of any gene or its variants across the continuous cardiometabolic spectrum from health (e.g. blood pressure traits) to disease (e.g. hypertension), facilitating the understanding of CVD trait genetics that inform health-to-disease transitions and endophenotypes. Custom visualizations in the new and improved user interface, including performance enhancements and new security features such as user authentication, collectively re-imagine HeartBioPortal's user experience and provide a data commons that co-locates data, storage and computing infrastructure in the context of studying the genetic basis behind the leading cause of global mortality. Database URL: https://www.heartbioportal.com/.
© The Author(s) 2020. Published by Oxford University Press.
Figures
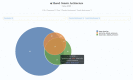
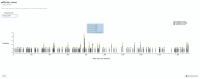


References
-
- Otto C.M., Savla J.J. and Hisama F.M. (2020) Cardiogenetics: a primer for the clinical cardiologist. Heart, 106, 938–947. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

