A glimpse of antimicrobial resistance gene diversity in kefir and yoghurt
- PMID: 33384459
- PMCID: PMC7775456
- DOI: 10.1038/s41598-020-80444-5
A glimpse of antimicrobial resistance gene diversity in kefir and yoghurt
Abstract
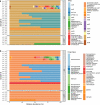
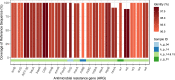
Antimicrobial resistance (AMR) is a global threat gaining more and more practical significance every year. The main determinants of AMR are the antimicrobial resistance genes (ARGs). Since bacteria can share genetic components via horizontal gene transfer, even non-pathogenic bacteria may provide ARG to any pathogens which they become physically close to (e.g. in the human gut). In addition, fermented food naturally contains bacteria in high amounts. In this study, we examined the diversity of ARG content in various kefir and yoghurt samples (products, grains, bacterial strains) using a unified metagenomic approach. We found numerous ARGs of commonly used fermenting bacteria. Even with the strictest filter restrictions, we identified ARGs undermining the efficacy of aminocoumarins, aminoglycosides, carbapenems, cephalosporins, cephamycins, diaminopyrimidines, elfamycins, fluoroquinolones, fosfomycins, glycylcyclines, lincosamides, macrolides, monobactams, nitrofurans, nitroimidazoles, penams, penems, peptides, phenicols, rifamycins, tetracyclines and triclosan. In the case of gene lmrD, we detected genetic environment providing mobility of this ARG. Our findings support the theory that during the fermentation process, the ARG content of foods can grow due to bacterial multiplication. The results presented suggest that the starting culture strains of fermented foods should be monitored and selected in order to decrease the intake of ARGs via foods.
Conflict of interest statement
The authors declare no competing interests.
Figures




References
-
- Machado M, et al. Antibiotic resistance and biofilm production in catalase-positive gram-positive cocci isolated from brazilian pasteurized milk. J. Food Qual. Hazards Control. 2020;7:67–74.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

