HOXA5 Expression Is Elevated in Breast Cancer and Is Transcriptionally Regulated by Estradiol
- PMID: 33384715
- PMCID: PMC7770181
- DOI: 10.3389/fgene.2020.592436
HOXA5 Expression Is Elevated in Breast Cancer and Is Transcriptionally Regulated by Estradiol
Abstract
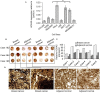
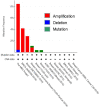
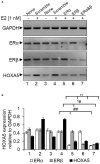
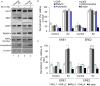
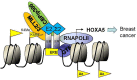
HOXA5 is a homeobox-containing gene associated with the development of the lung, gastrointestinal tract, and vertebrae. Here, we investigate potential roles and the gene regulatory mechanism in HOXA5 in breast cancer cells. Our studies demonstrate that HOXA5 expression is elevated in breast cancer tissues and in estrogen receptor (ER)-positive breast cancer cells. HOXA5 expression is critical for breast cancer cell viability. Biochemical studies show that estradiol (E2) regulates HOXA5 gene expression in cultured breast cancer cells in vitro. HOXA5 expression is also upregulated in vivo in the mammary tissues of ovariectomized female rats. E2-induced HOXA5 expression is coordinated by ERs. Knockdown of either ERα or ERβ downregulated E2-induced HOXA5 expression. Additionally, ER co-regulators, including CBP/p300 (histone acetylases) and MLL-histone methylases (MLL2, MLL3), histone acetylation-, and H3K4 trimethylation levels are enriched at the HOXA5 promoter in present E2. In summary, our studies demonstrate that HOXA5 is overexpressed in breast cancer and is transcriptionally regulated via estradiol in breast cancer cells.
Keywords: HOXA5; breast cancer; chromatin; estradiol; estrogen receptors; gene expression.
Copyright © 2020 Hussain, Deb, Chini, Obaid, Bhan, Ansari, Mishra, Bobzean, Udden, Alluri, Das, Brothers, Perrotti and Mandal.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Antisense transcript long noncoding RNA (lncRNA) HOTAIR is transcriptionally induced by estradiol.J Mol Biol. 2013 Oct 9;425(19):3707-22. doi: 10.1016/j.jmb.2013.01.022. Epub 2013 Jan 31. J Mol Biol. 2013. PMID: 23375982 Free PMC article.
-
HOXC6 Is transcriptionally regulated via coordination of MLL histone methylase and estrogen receptor in an estrogen environment.J Mol Biol. 2011 Aug 12;411(2):334-49. doi: 10.1016/j.jmb.2011.05.050. Epub 2011 Jun 12. J Mol Biol. 2011. PMID: 21683083 Free PMC article.
-
Bisphenol-A induces expression of HOXC6, an estrogen-regulated homeobox-containing gene associated with breast cancer.Biochim Biophys Acta. 2015 Jun;1849(6):697-708. doi: 10.1016/j.bbagrm.2015.02.003. Epub 2015 Feb 25. Biochim Biophys Acta. 2015. PMID: 25725483 Free PMC article.
-
Endocrine disrupting chemical, bisphenol-A, induces breast cancer associated gene HOXB9 expression in vitro and in vivo.Gene. 2016 Sep 30;590(2):234-43. doi: 10.1016/j.gene.2016.05.009. Epub 2016 May 13. Gene. 2016. PMID: 27182052 Free PMC article.
-
Hoxa5: A Key Player in Development and Disease.J Dev Biol. 2016 Mar 25;4(2):13. doi: 10.3390/jdb4020013. J Dev Biol. 2016. PMID: 29615582 Free PMC article. Review.
Cited by
-
The hidden messengers: cancer associated fibroblasts-derived exosomal miRNAs as key regulators of cancer malignancy.Front Cell Dev Biol. 2024 Apr 17;12:1378302. doi: 10.3389/fcell.2024.1378302. eCollection 2024. Front Cell Dev Biol. 2024. PMID: 38694824 Free PMC article. Review.
-
Novel long non-coding RNAs associated with inflammation and macrophage activation in human.Sci Rep. 2023 Mar 10;13(1):4036. doi: 10.1038/s41598-023-30568-1. Sci Rep. 2023. PMID: 36899011 Free PMC article.
-
The KDM5A/HOXA5 axis regulates osteosarcoma progression via activating the Wnt/β-catenin pathway.Eur J Med Res. 2025 Apr 15;30(1):284. doi: 10.1186/s40001-025-02478-7. Eur J Med Res. 2025. PMID: 40229896 Free PMC article.
-
Expression Landscape and Functional Roles of HOXA4 and HOXA5 in Lung Adenocarcinoma.Int J Med Sci. 2022 Mar 6;19(3):572-587. doi: 10.7150/ijms.70445. eCollection 2022. Int J Med Sci. 2022. PMID: 35370463 Free PMC article.
-
Cribriform morular thyroid carcinoma: Clinicopathological and molecular basis for both a preventive and therapeutic approach for a rare tumor (Review).Oncol Rep. 2024 Sep;52(3):119. doi: 10.3892/or.2024.8778. Epub 2024 Jul 19. Oncol Rep. 2024. PMID: 39027989 Free PMC article. Review.
References
-
- Akam M. (1987). The molecular basis for metameric pattern in the Drosophila embryo. Development 101, 1–22. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

