Genomic Epidemiology of MRSA During Incarceration at a Large Inner-City Jail
- PMID: 33395473
- PMCID: PMC8664465
- DOI: 10.1093/cid/ciaa1937
Genomic Epidemiology of MRSA During Incarceration at a Large Inner-City Jail
Abstract
Background: Congregate settings, such as jails, may be a location where colonized detainees transmit methicillin-resistant Staphylococcus aureus (MRSA). We examined MRSA acquisition during incarceration and characterized the genomic epidemiology of MRSA entering the jail and isolated during incarceration.
Methods: Males incarcerated at the Cook County Jail were enrolled within 72 h of intake and MRSA surveillance cultures collected. Detainees in jail at Day 30 were re-cultured to determine MRSA acquisition. A survey was administered to identify acquisition predictors. Genomic sequencing of surveillance and clinical isolates was integrated with epidemiologic and jail location data to track MRSA transmission pathways.
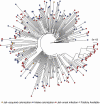
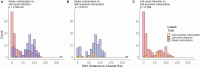
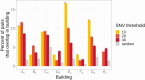
Results: 800 males were enrolled; 19% MRSA colonized at intake. Of 184 who reached Day 30 visit, 12 acquired MRSA. Heroin use before entering (OR 3.67, P = .05) and sharing personal items during incarceration (OR = 4.92, P = .01) were predictors of acquisition. Sequenced clinical USA300 isolates (n = 112) were more genetically similar than diverse intake USA300 strains (P < .001), suggesting jail transmission. Four acquired colonization isolates were within 20 single-nucleotide variant (SNVs) of other isolates; 4 were within 20 SNVs of an intake isolate, 2 for an acquisition isolate, and 1 for a clinical isolate. Individuals with genetically similar isolates were more likely to have had overlapping stays in the same buildings.
Conclusion: There was a high MRSA burden entering jail. Genomic analysis of acquisition and clinical isolates suggests potential spread of incoming strains and networks of spread during incarceration, with spread often occurring among detainees housed in similar locations. Sharing personal items during incarceration is associated with MRSA acquisition and could be a focus for intervention.
Keywords: MRSA; Whole genome sequencing; jail.
© The Author(s) 2021. Published by Oxford University Press for the Infectious Diseases Society of America. All rights reserved. For permissions, e-mail: journals.permissions@oup.com.
Figures

References
-
- Fridkin SK, Hageman JC, Morrison M, et al. ; Active Bacterial Core Surveillance Program of the Emerging Infections Program Network . Methicillin-resistant Staphylococcus aureus disease in three communities. N Engl J Med 2005; 352:1436–44. - PubMed
-
- Miller LG, Diep BA. Clinical practice: colonization, fomites, and virulence: rethinking the pathogenesis of community-associated methicillin-resistant Staphylococcus aureus infection. Clin Infect Dis 2008; 46:752–60. - PubMed
-
- Popovich KJ, Hota B, Weinstein RA. Treatment of community-associated methicillin-resistant Staphylococcus aureus. Curr Infect Dis Rep 2007; 9:398–407. - PubMed
-
- Methicillin-resistant Staphylococcus aureus infections in correctional facilities–Georgia, California, and Texas, 2001–2003. MMWR Morb Mortal Wkly Rep 2003; 52:992–6. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

