Molecular and Biochemical Techniques for Deciphering p53-MDM2 Regulatory Mechanisms
- PMID: 33396576
- PMCID: PMC7824699
- DOI: 10.3390/biom11010036
Molecular and Biochemical Techniques for Deciphering p53-MDM2 Regulatory Mechanisms
Abstract
The p53 and Mouse double minute 2 (MDM2) proteins are hubs in extensive networks of interactions with multiple partners and functions. Intrinsically disordered regions help to adopt function-specific structural conformations in response to ligand binding and post-translational modifications. Different techniques have been used to dissect interactions of the p53-MDM2 pathway, in vitro, in vivo, and in situ each having its own advantages and disadvantages. This review uses the p53-MDM2 to show how different techniques can be employed, illustrating how a combination of in vitro and in vivo techniques is highly recommended to study the spatio-temporal location and dynamics of interactions, and to address their regulation mechanisms and functions. By using well-established techniques in combination with more recent advances, it is possible to rapidly decipher complex mechanisms, such as the p53 regulatory pathway, and to demonstrate how protein and nucleotide ligands in combination with post-translational modifications, result in inter-allosteric and intra-allosteric interactions that govern the activity of the protein complexes and their specific roles in oncogenesis. This promotes elegant therapeutic strategies that exploit protein dynamics to target specific interactions.
Keywords: ATM; DNA damage response; MDM2; MDMX; p53; p53 mRNA; post-translational modification; protein-RNA interactions; protein-protein interactions.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
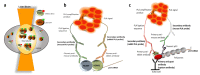

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

