The interplay between emerging human coronavirus infections and autophagy
- PMID: 33399028
- PMCID: PMC7872537
- DOI: 10.1080/22221751.2021.1872353
The interplay between emerging human coronavirus infections and autophagy
Abstract
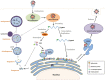
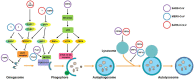
ABSTRACT Following outbreaks of severe acute respiratory syndrome coronavirus (SARS-CoV) and the Middle East respiratory syndrome coronavirus (MERS-CoV) in 2002 and 2012, respectively, the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the third highly pathogenic emerging human coronavirus (hCoV). SARS-CoV-2 is currently causing the global coronavirus disease 2019 (COVID-19) pandemic. CoV infections in target cells may stimulate the formation of numerous double-membrane autophagosomes and induce autophagy. Several studies provided evidence that hCoV infections are closely related to various cellular aspects associated with autophagy. Autophagy may even promote hCoV infection and replication. However, so far it is unclear how hCoV infections induce autophagy and whether the autophagic machinery is necessary for viral propagation. Here, we summarize the most recent advances concerning the mutual interplay between the autophagic machinery and the three emerging hCoVs, SARS-CoV, MERS-CoV, and SARS-CoV-2 and the model system mouse hepatitis virus. We also discuss the applicability of approved and well-tolerated drugs targeting autophagy as a potential treatment against COVID-19.
Keywords: Autophagy; MERS-CoV; MHV; SARS-CoV; SARS-CoV-2; coronavirus.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures
References
-
- Drosten C, Gunther S, Preiser W, et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N Engl J Med. 2003;348:1967–1976. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous
