Construction and application of a human scFv phage display library based on Cre‑LoxP recombination for anti‑PCSK9 antibody selection
- PMID: 33416098
- PMCID: PMC7797424
- DOI: 10.3892/ijmm.2020.4822
Construction and application of a human scFv phage display library based on Cre‑LoxP recombination for anti‑PCSK9 antibody selection
Abstract
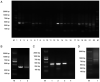
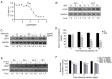
A large human natural single‑chain fragment variable (scFv) phage library was constructed based on Cre‑LoxP recombination, and used to successfully identify antibodies against proprotein convertase subtilisin/kexin type 9 (PCSK9). The library was derived from 400 blood samples, 30 bone marrow samples, and 10 cord blood samples from healthy donors. Lymphocytes were isolated from each sample and cDNA was synthesized using reverse transcription‑quantitative PCR. Two‑step overlap PCR was then used for scFv synthesis using a LoxP peptide as the linker. The scFv gene was inserted into the phagemid vector pDF by enzymatic digestion and ligation, and then transformed into Escherichia coli (E. coli) SS320 to establish a primary antibody library in the form of scFvs. A primary antibody library consisting of 5x107 peripheral blood and umbilical cord blood sources, as well as a primary antibody library of 5x107 bone marrow samples were obtained. By optimizing the recombination conditions, the primary phage library was used to infect E. coli BS1365 strain (which expresses the Cre enzyme), and a human scFv recombinant library with a size of 1x1011 was obtained through Cre‑LoxP enzyme‑mediated heavy and light chain replacement and recombination. This constructed recombinant library was employed to screen for antibodies against recombinant PCSK9. After four rounds of selection, a fully human antibody (3D2) was identified with a binding affinity of 1.96±1.56ⅹ10‑10 M towards PCSK9. In vitro, the PCSK9/low‑density lipoprotein receptor (LDLR) pathway of Hep‑G2 cells was inhibited by 3D2 treatment, thereby increasing LDL uptake in these cells. In addition, combination treatment with 3D2 and statin was more effective at increasing LDLR levels than treatment with 3D2 or statin alone. Furthermore, 3D2 resulted in a 3‑fold increase in hepatic LDLR levels, and lowered total serum cholesterol by up to 61.5% in vivo. Taken together, these results suggest that the constructed human Cre‑LoxP scFv phage display library can be used to screen fully human scFv, and that 3D2 may serve as a candidate hypolipidemic therapy.
Keywords: proprotein convertase subtilisin-like/kexin type 9; low-density lipoprotein; receptor; fully human single-chain fragment variable phage display antibody library; Cre-LoxP; hypercholesterolemia.
Figures




References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

