Tissue-resident CD4+ T helper cells assist the development of protective respiratory B and CD8+ T cell memory responses
- PMID: 33419791
- PMCID: PMC8056937
- DOI: 10.1126/sciimmunol.abb6852
Tissue-resident CD4+ T helper cells assist the development of protective respiratory B and CD8+ T cell memory responses
Abstract
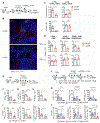
Much remains unknown about the roles of CD4+ T helper cells in shaping localized memory B cell and CD8+ T cell immunity in the mucosal tissues. Here, we report that lung T helper cells provide local assistance for the optimal development of tissue-resident memory B and CD8+ T cells after the resolution of primary influenza virus infection. We have identified a population of T cells in the lung that exhibit characteristics of both follicular T helper and TRM cells, and we have termed these cells as resident helper T (TRH) cells. Optimal TRH cell formation was dependent on transcription factors involved in T follicular helper and resident memory T cell development including BCL6 and Bhlhe40. We show that TRH cells deliver local help to CD8+ T cells through IL-21-dependent mechanisms. Our data have uncovered the presence of a tissue-resident helper T cell population in the lung that plays a critical role in promoting the development of protective B cell and CD8+ T cell responses.
Copyright © 2021 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Conflict of interest statement
Competing interests: The authors declare that they have no competing interests.
Figures






Comment in
-
TRH cells, helpers making an impact in their local community.Sci Immunol. 2021 Jan 8;6(55):eabf2886. doi: 10.1126/sciimmunol.abf2886. Sci Immunol. 2021. PMID: 33419792
References
-
- Mueller SN, Mackay LK, Tissue-resident memory T cells: Local specialists in immune defence. Nat. Rev. Immunol. 16, 79–89 (2016). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials