Thioesterase-mediated side chain transesterification generates potent Gq signaling inhibitor FR900359
- PMID: 33420046
- PMCID: PMC7794379
- DOI: 10.1038/s41467-020-20418-3
Thioesterase-mediated side chain transesterification generates potent Gq signaling inhibitor FR900359
Abstract
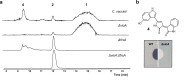
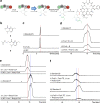
The potent and selective Gq protein inhibitor depsipeptide FR900359 (FR), originally discovered as the product of an uncultivable plant endosymbiont, is synthesized by a complex biosynthetic system comprising two nonribosomal peptide synthetase (NRPS) assembly lines. Here we characterize a cultivable bacterial FR producer, enabling detailed investigations into biosynthesis and attachment of the functionally important FR side chain. We reconstitute side chain assembly by the monomodular NRPS FrsA and the non-heme monooxygenase FrsH, and characterize intermolecular side chain transesterification to the final macrocyclic intermediate FR-Core, mediated by the FrsA thioesterase domain. We harness FrsA substrate promiscuity to generate FR analogs with altered side chains and demonstrate indispensability of the FR side chain for efficient Gq inhibition by comparative bioactivity, toxicity and docking studies. Finally, evolution of FR and side chain biosynthesis is discussed based on bioinformatics analyses. Side chain transesterification boosts potency and target affinity of selective Gq inhibitor natural products.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Heterologous Expression, Biosynthetic Studies, and Ecological Function of the Selective Gq-Signaling Inhibitor FR900359.Angew Chem Int Ed Engl. 2018 Jan 15;57(3):836-840. doi: 10.1002/anie.201707996. Epub 2017 Dec 20. Angew Chem Int Ed Engl. 2018. PMID: 29194875
-
Promoter-Driven Overexpression in Chromobacterium vaccinii Facilitates Access to FR900359 and Yields Novel Low Abundance Analogs.Chemistry. 2022 Feb 7;28(8):e202103888. doi: 10.1002/chem.202103888. Epub 2021 Dec 28. Chemistry. 2022. PMID: 34878202
-
Feature-Based Molecular Networking for the Targeted Identification of Gq-Inhibiting FR900359 Derivatives.J Nat Prod. 2021 Jul 23;84(7):1941-1953. doi: 10.1021/acs.jnatprod.1c00194. Epub 2021 Jul 1. J Nat Prod. 2021. PMID: 34197116
-
The chromodepsins - chemistry, biology and biosynthesis of a selective Gq inhibitor natural product family.Nat Prod Rep. 2021 Dec 15;38(12):2276-2292. doi: 10.1039/d1np00005e. Nat Prod Rep. 2021. PMID: 33998635 Review.
-
Heterotrimeric Gq proteins as therapeutic targets?J Biol Chem. 2020 Apr 17;295(16):5206-5215. doi: 10.1074/jbc.REV119.007061. Epub 2020 Mar 2. J Biol Chem. 2020. PMID: 32122969 Free PMC article. Review.
Cited by
-
Macrocyclic Gq Protein Inhibitors FR900359 and/or YM-254890-Fit for Translation?ACS Pharmacol Transl Sci. 2021 Feb 19;4(2):888-897. doi: 10.1021/acsptsci.1c00021. eCollection 2021 Apr 9. ACS Pharmacol Transl Sci. 2021. PMID: 33860209 Free PMC article.
-
Plant peptides - redefining an area of ribosomally synthesized and post-translationally modified peptides.Nat Prod Rep. 2024 Jul 17;41(7):1020-1059. doi: 10.1039/d3np00042g. Nat Prod Rep. 2024. PMID: 38411572 Free PMC article. Review.
-
Structural response of G protein binding to the cyclodepsipeptide inhibitor FR900359 probed by NMR spectroscopy.Chem Sci. 2024 Jul 4;15(32):12939-12956. doi: 10.1039/d4sc01950d. eCollection 2024 Aug 14. Chem Sci. 2024. PMID: 39148790 Free PMC article.
-
A Specialized Dehydrogenase Provides l-Phenyllactate for FR900359 Biosynthesis.Chembiochem. 2022 May 18;23(10):e202100569. doi: 10.1002/cbic.202100569. Epub 2021 Dec 9. Chembiochem. 2022. PMID: 34846772 Free PMC article.
-
Recommended Tool Compounds: Application of YM-254890 and FR900359 to Interrogate Gαq/11-Mediated Signaling Pathways.ACS Pharmacol Transl Sci. 2023 Nov 10;6(12):1790-1800. doi: 10.1021/acsptsci.3c00214. eCollection 2023 Dec 8. ACS Pharmacol Transl Sci. 2023. PMID: 38093837 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources

