HIV-1 diversity considerations in the application of the Intact Proviral DNA Assay (IPDA)
- PMID: 33420062
- PMCID: PMC7794580
- DOI: 10.1038/s41467-020-20442-3
HIV-1 diversity considerations in the application of the Intact Proviral DNA Assay (IPDA)
Erratum in
-
Author Correction: HIV-1 diversity considerations in the application of the Intact Proviral DNA Assay (IPDA).Nat Commun. 2021 May 13;12(1):2958. doi: 10.1038/s41467-021-23515-z. Nat Commun. 2021. PMID: 33986282 Free PMC article. No abstract available.
Abstract
The Intact Proviral DNA Assay (IPDA) was developed to address the critical need for a scalable method for intact HIV-1 reservoir quantification. This droplet digital PCR-based assay simultaneously targets two HIV-1 regions to distinguish genomically intact proviruses against a large background of defective ones, and its application has yielded insights into HIV-1 persistence. Reports of assay failures however, attributed to HIV-1 polymorphism, have recently emerged. Here, we describe a diverse North American cohort of people with HIV-1 subtype B, where the IPDA yielded a failure rate of 28% due to viral polymorphism. We further demonstrate that within-host HIV-1 diversity can lead the IPDA to underestimate intact reservoir size, and provide examples of how this phenomenon could lead to erroneous interpretation of clinical trial data. While the IPDA represents a major methodological advance, HIV-1 diversity should be addressed before its widespread adoption as a principal readout in HIV-1 remission trials.
Conflict of interest statement
The authors declare no competing interests.
Figures


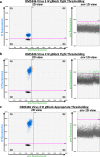

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

