Genetic Variation, GWAS and Accuracy of Prediction for Host Resistance to Sparicotyle chrysophrii in Farmed Gilthead Sea Bream (Sparus aurata)
- PMID: 33424925
- PMCID: PMC7793675
- DOI: 10.3389/fgene.2020.594770
Genetic Variation, GWAS and Accuracy of Prediction for Host Resistance to Sparicotyle chrysophrii in Farmed Gilthead Sea Bream (Sparus aurata)
Abstract
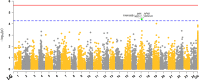
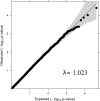
Gilthead sea bream (Sparus aurata) belongs to a group of teleost which has high importance in Mediterranean aquaculture industry. However, industrial production is increasingly compromised by an elevated outbreak of diseases in sea cages, especially a disease caused by monogeneans parasite Sparicotyle chrysophrii. This parasite mainly colonizes gill tissues of host and causes considerable economical losses with mortality and reduction in growth. The aim of current study was to explore the genetics of host resistance against S. chrysophrii and investigate the potential for genomic selection to possibly accelerate genetic progress. To achieve the desired goals, a test population derived from the breeding nucleus of Andromeda Group was produced. This experimental population was established by crossing of parents mated in partial factorial crosses of ∼8 × 8 using 58 sires and 62 dams. The progeny obtained from this mating design was challenged with S. chrysophrii using a controllable cohabitation infection model. At the end of the challenge, fish were recorded for parasite count, and all the recorded fish were tissue sampled for genotyping by sequencing using 2b-RAD methodology. The initial (before challenge test) and the final body weight (after challenge test) of the fish were also recorded. The results obtained through the analysis of phenotypic records (n = 615) and the genotypic data (n = 841, 724 offspring and 117 parents) revealed that the resistance against this parasite is lowly heritable (h 2 = 0.147 with pedigree and 0.137 with genomic information). We observed moderately favorable genetic correlation (R g = -0.549 to -0.807) between production traits (i.e., body weight and specific growth rate) and parasite count, which signals a possibility of indirect selection. A locus at linkage group 17 was identified that surpassed chromosome-wide Bonferroni threshold which explained 22.68% of the total genetic variance, and might be playing role in producing genetic variation. The accuracy of prediction was improved by 8% with genomic information compared to pedigree.
Keywords: 2b-RAD sequencing; genetic correlation; genetic variation; genome-wide association analysis; genomic selection; single nucleotide polymorphisms.
Copyright © 2020 Aslam, Carraro, Sonesson, Meuwissen, Tsigenopoulos, Rigos, Bargelloni and Tzokas.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Genetic variation of Sparicotyle chrysophrii (Monogenea: Microcotylidae) from the gilthead sea bream Sparus aurata (Teleostei: Sparidae) in the Mediterranean Sea.Parasitol Res. 2023 Jan;122(1):157-165. doi: 10.1007/s00436-022-07709-y. Epub 2022 Nov 24. Parasitol Res. 2023. PMID: 36418649
-
Genetics of resistance to photobacteriosis in gilthead sea bream (Sparus aurata) using 2b-RAD sequencing.BMC Genet. 2018 Jul 11;19(1):43. doi: 10.1186/s12863-018-0631-x. BMC Genet. 2018. PMID: 29996763 Free PMC article.
-
Genomic Prediction of Resistance to Pasteurellosis in Gilthead Sea Bream (Sparus aurata) Using 2b-RAD Sequencing.G3 (Bethesda). 2016 Nov 8;6(11):3693-3700. doi: 10.1534/g3.116.035220. G3 (Bethesda). 2016. PMID: 27652890 Free PMC article.
-
Acting locally - affecting globally: RNA sequencing of gilthead sea bream with a mild Sparicotyle chrysophrii infection reveals effects on apoptosis, immune and hypoxia related genes.BMC Genomics. 2019 Mar 11;20(1):200. doi: 10.1186/s12864-019-5581-9. BMC Genomics. 2019. PMID: 30866816 Free PMC article.
-
Mapping the knowledge of the main diseases affecting sea bass and sea bream in Mediterranean.Transbound Emerg Dis. 2020 May;67(3):1089-1100. doi: 10.1111/tbed.13482. Epub 2020 Jan 30. Transbound Emerg Dis. 2020. PMID: 31960605 Review.
Cited by
-
The first high-density genetic map of common cockle (Cerastoderma edule) reveals a major QTL controlling shell color variation.Sci Rep. 2022 Oct 10;12(1):16971. doi: 10.1038/s41598-022-21214-3. Sci Rep. 2022. PMID: 36216849 Free PMC article.
-
Applying genetic technologies to combat infectious diseases in aquaculture.Rev Aquac. 2023 Mar;15(2):491-535. doi: 10.1111/raq.12733. Epub 2022 Sep 5. Rev Aquac. 2023. PMID: 38504717 Free PMC article. Review.
-
Genetic Basis for Resistance Against Viral Nervous Necrosis: GWAS and Potential of Genomic Prediction Explored in Farmed European Sea Bass (Dicentrarchus labrax).Front Genet. 2022 Mar 25;13:804584. doi: 10.3389/fgene.2022.804584. eCollection 2022. Front Genet. 2022. PMID: 35401661 Free PMC article.
-
The effect of cycles of genomic selection on the wheat (T. aestivum) genome.Theor Appl Genet. 2023 Mar 23;136(4):70. doi: 10.1007/s00122-023-04279-0. Theor Appl Genet. 2023. PMID: 36952091
-
Mucosal affairs: glycosylation and expression changes of gill goblet cells and mucins in a fish-polyopisthocotylidan interaction.Front Vet Sci. 2024 Apr 9;11:1347707. doi: 10.3389/fvets.2024.1347707. eCollection 2024. Front Vet Sci. 2024. PMID: 38655531 Free PMC article.
References
-
- Aslam M. L., Carraro R., Sonesson A., Tzokas K., Tsigenopoulos C., Rigos G., et al. (2018b). “Genetic basis of host resistance to S. chrysophrii in farmed gilthead sea bream (Sparus aurata) population”, Proceedings of the World Congress on Genetics Applied to Livestock Production, 2:39.
LinkOut - more resources
Full Text Sources

