Population structure of indigenous inhabitants of Arabia
- PMID: 33428619
- PMCID: PMC7799765
- DOI: 10.1371/journal.pgen.1009210
Population structure of indigenous inhabitants of Arabia
Abstract
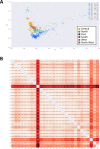
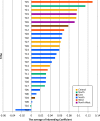
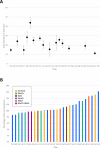
Modern day Saudi Arabia occupies the majority of historical Arabia, which may have contributed to ancient waves of migration out of Africa. This ancient history has left a lasting imprint in the genetics of the region, including the diverse set of tribes that call Saudi Arabia their home. How these tribes relate to each other and to the world's major populations remains an unanswered question. In an attempt to improve our understanding of the population structure of Saudi Arabia, we conducted genomic profiling of 957 unrelated individuals who self-identify with 28 large tribes in Saudi Arabia. Consistent with the tradition of intra-tribal unions, the subjects showed strong clustering along tribal lines with the distance between clusters correlating with their geographical proximities in Arabia. However, these individuals form a unique cluster when compared to the world's major populations. The ancient origin of these tribal affiliations is supported by analyses that revealed little evidence of ancestral origin from within the 28 tribes. Our results disclose a granular map of population structure and have important implications for future genetic studies into Mendelian and common diseases in the region.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

