Epigenomic Profiles of African-American Transthyretin Val122Ile Carriers Reveals Putatively Dysregulated Amyloid Mechanisms
- PMID: 33428857
- PMCID: PMC7887108
- DOI: 10.1161/CIRCGEN.120.003011
Epigenomic Profiles of African-American Transthyretin Val122Ile Carriers Reveals Putatively Dysregulated Amyloid Mechanisms
Abstract
Background: The Val122Ile mutation in Transthyretin (TTR) gene causes a rare, difficult to diagnose hereditary form of cardiac amyloidosis. This mutation is most common in the United States and mainly present in people of African descent. The carriers have an increased risk of congestive heart failure, peripheral edema, and several other noncardiac phenotypes such as carpal tunnel syndrome, and arthroplasty which are top reasons for ambulatory/outpatient surgeries (OSs) in the country.
Methods: We conducted first-ever epigenome-wide association study using the Illumina's EPIC array, in Val122Ile carriers of African descent for heart disease and multiple OSs-an early disease indicator. Differential methylation across genome wide cytosine-phosphate guanine (CpG) sites was tested between carriers with and without heart disease and OS. Significant CpG sites were investigated for cis-mQTLs loci, followed by gene ontology and protein-protein interaction network. We also investigated the significant CpG sites in a secondary cohort of carriers for replication.
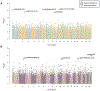
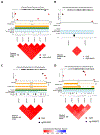
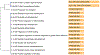
Results: Five differentially methylated sites (P≤2.1×10-8) in genes-FAM129B, SKI, WDR27, GLS, and an intergenic site near RP11-550A5.2, and one differentially methylated region containing KCNA6 and GALNT3 (P=1.1×10-12) were associated with heart disease. For OS, we observe 4 sites-2 sites in UBE2E3 and SEC14L5, and other 2 in intergenic regions (P≤1.8×10-7) and 3 regions overlapping SH3D21, EVA1B, LTB4R2, and CIDEB (P≤3.9×10-7). Functional protein-interaction module analysis identified ABCA1 (P=0.001) for heart disease. Six cis-mQTLs were associated with one of the significant CpG sites (FAM129B; P=4.1×10-24). We replicated 2 CpG sites (cg18546846 and cg06641417; P<0.05) in an external cohort of biopsy-confirmed cases of TTR (transthyretin) amyloidosis. The genes identified are involved in transport and clearance of amyloid deposits (GLS, ABCA1, FAM129B); cardiac fibrosis (SKI); and muscle tissue regulation (SKI, FAM129B).
Conclusions: These findings highlight the link between a complex amyloid circuit and diverse symptoms of Val122Ile.
Keywords: amyloid; edema; methylation; mutation.
Figures



Similar articles
-
Association of Transthyretin Val122Ile Variant With Incident Heart Failure Among Black Individuals.JAMA. 2022 Apr 12;327(14):1368-1378. doi: 10.1001/jama.2022.2896. JAMA. 2022. PMID: 35377943 Free PMC article.
-
Epigenetic profiling of Italian patients identified methylation sites associated with hereditary transthyretin amyloidosis.Clin Epigenetics. 2020 Nov 17;12(1):176. doi: 10.1186/s13148-020-00967-6. Clin Epigenetics. 2020. PMID: 33203445 Free PMC article.
-
DISCOVERY: prevalence of transthyretin (TTR) mutations in a US-centric patient population suspected of having cardiac amyloidosis.Amyloid. 2020 Dec;27(4):223-230. doi: 10.1080/13506129.2020.1764928. Epub 2020 May 26. Amyloid. 2020. PMID: 32456532
-
Transthyretin V122I (pV142I)* cardiac amyloidosis: an age-dependent autosomal dominant cardiomyopathy too common to be overlooked as a cause of significant heart disease in elderly African Americans.Genet Med. 2017 Jul;19(7):733-742. doi: 10.1038/gim.2016.200. Epub 2017 Jan 19. Genet Med. 2017. PMID: 28102864 Free PMC article. Review.
-
Cardiac Amyloidosis Due to Transthyretin Protein: A Review.JAMA. 2024 Mar 5;331(9):778-791. doi: 10.1001/jama.2024.0442. JAMA. 2024. PMID: 38441582 Free PMC article. Review.
Cited by
-
Clinical spectrum of Transthyretin amyloidogenic mutations among diverse population origins.Hum Genomics. 2024 Mar 25;18(1):31. doi: 10.1186/s40246-024-00596-7. Hum Genomics. 2024. PMID: 38523305 Free PMC article.
-
Deep-embedded clustering by relevant scales and genome-wide association study in autism.PLoS One. 2025 May 29;20(5):e0322698. doi: 10.1371/journal.pone.0322698. eCollection 2025. PLoS One. 2025. PMID: 40440618 Free PMC article.
-
The integration of genetically-regulated transcriptomics and electronic health records highlights a pattern of medical outcomes related to increased hepatic transthyretin expression.Amyloid. 2022 Jun;29(2):110-119. doi: 10.1080/13506129.2021.2018678. Epub 2021 Dec 22. Amyloid. 2022. PMID: 34935565 Free PMC article.
-
Associations between pathophysiological traits and symptom development in retrospective analysis of V30M and V122I transthyretin amyloidosis.Int J Cardiol Heart Vasc. 2025 Apr 15;58:101663. doi: 10.1016/j.ijcha.2025.101663. eCollection 2025 Jun. Int J Cardiol Heart Vasc. 2025. PMID: 40276302 Free PMC article.
-
Hereditary transthyretin amyloidosis: a myriad of factors that influence phenotypic variability.J Neurol. 2024 Sep;271(9):5746-5761. doi: 10.1007/s00415-024-12509-8. Epub 2024 Jun 22. J Neurol. 2024. PMID: 38907862 Free PMC article. Review.
References
-
- Benson MD, Buxbaum JN, Eisenberg DS, Merlini G, Saraiva MJM, Sekijima Y, Sipe JD, Westermark P. Amyloid nomenclature 2018: recommendations by the International Society of Amyloidosis (ISA) nomenclature committee. Amyloid. 2018;25:215–219. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

