Broad-range and effective detection of human noroviruses by colloidal gold immunochromatographic assay based on the shell domain of the major capsid protein
- PMID: 33430771
- PMCID: PMC7798207
- DOI: 10.1186/s12866-020-02084-z
Broad-range and effective detection of human noroviruses by colloidal gold immunochromatographic assay based on the shell domain of the major capsid protein
Abstract
Background: Human noroviruses (HuNoVs) are a major cause of nonbacterial gastroenteritis in all age groups worldwide. HuNoVs can be detected in vitro using molecular assays such as RT-PCR and RT-qPCR. However, these molecular-based techniques require special equipment, unique reagents, experienced personnel, and extended time to obtain results. Besides, the diversity of viral genotypes is high. Therefore, methods that are rapid, broad-range and effective in the detection of HuNoVs are desiderated for screening the feces or vomit of infected people during outbreaks.
Results: In this study, a colloidal-gold-based immunochromatographic assay (ICA) was developed for effective detection of HuNoVs in clinical samples. Monoclonal antibodies (MAbs) against the shell (S) domain in the major capsid protein of HuNoVs were used in the ICA. The limitations of detection for HuNoVs in clinical samples were 1.2 × 106 genomic copies per gram of stool sample (gc/g) and 4.4 × 105 gc/g for genogroup I and II (GI and GII) HuNoVs, respectively. A total of 122 clinical samples were tested for HuNoVs by ICA and compared against RT-qPCR. The relative sensitivity, specificity and agreement of ICA was 84.2% (95% CI: 83.6-84.8%), 100.0% (95% CI: 98.5-100.0%) and 87.7% (95% CI: 85.6-89.8%), respectively. No cross-reaction with other common enteric viruses or bacteria was observed. The ICA detected a broad range of genotypes, including GI.1, GI.3, GI.4, GI.6, GI.14, GII.2, GII.3, GII.4, GII.6, GII.13, and GII.17 HuNoVs.
Conclusions: This study demonstrates that ICA targeting the S domain of VP1 is a promising candidate for effectively identifying the different genotypes of HuNoVs in clinical samples with high sensitivity and specificity.
Keywords: Broad-range detection; Colloid gold; Human norovirus; Immunochromatographic assays; S domain of VP1.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


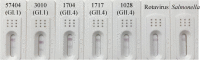
Similar articles
-
Fingerprinting of human noroviruses co-infections in a possible foodborne outbreak by metagenomics.Int J Food Microbiol. 2020 Nov 16;333:108787. doi: 10.1016/j.ijfoodmicro.2020.108787. Epub 2020 Jul 9. Int J Food Microbiol. 2020. PMID: 32702583
-
Rapid detection of norovirus from fecal specimens by real-time reverse transcription-loop-mediated isothermal amplification assay.J Clin Microbiol. 2006 Apr;44(4):1376-81. doi: 10.1128/JCM.44.4.1376-1381.2006. J Clin Microbiol. 2006. PMID: 16597865 Free PMC article.
-
Evaluation of the Dako IDEIA norovirus EIA assay for detection of norovirus using faecal specimens from Australian gastroenteritis outbreaks.Pathology. 2006 Apr;38(2):157-65. doi: 10.1080/00313020600559645. Pathology. 2006. PMID: 16581657
-
The attachment factors and attachment receptors of human noroviruses.Food Microbiol. 2024 Oct;123:104591. doi: 10.1016/j.fm.2024.104591. Epub 2024 Jul 2. Food Microbiol. 2024. PMID: 39038896 Review.
-
Conformational Flexibility in Capsids Encoded by the Caliciviridae.Viruses. 2024 Nov 26;16(12):1835. doi: 10.3390/v16121835. Viruses. 2024. PMID: 39772145 Free PMC article. Review.
Cited by
-
Recent Advances in DNA Nanotechnology-Enabled Biosensors for Virus Detection.Biosensors (Basel). 2023 Aug 15;13(8):822. doi: 10.3390/bios13080822. Biosensors (Basel). 2023. PMID: 37622908 Free PMC article. Review.
-
Advances and Future Perspective on Detection Technology of Human Norovirus.Pathogens. 2021 Oct 26;10(11):1383. doi: 10.3390/pathogens10111383. Pathogens. 2021. PMID: 34832539 Free PMC article. Review.
-
The Road to Unconventional Detections: Paper-Based Microfluidic Chips.Micromachines (Basel). 2022 Oct 27;13(11):1835. doi: 10.3390/mi13111835. Micromachines (Basel). 2022. PMID: 36363856 Free PMC article. Review.
-
Ten Years of Lateral Flow Immunoassay Technique Applications: Trends, Challenges and Future Perspectives.Sensors (Basel). 2021 Jul 30;21(15):5185. doi: 10.3390/s21155185. Sensors (Basel). 2021. PMID: 34372422 Free PMC article. Review.
-
Comparative Analysis of RT-PCR and a Colloidal Gold Immunochromatographic Assay for SARS-CoV-2 Detection.Diagnostics (Basel). 2025 May 28;15(11):1362. doi: 10.3390/diagnostics15111362. Diagnostics (Basel). 2025. PMID: 40506934 Free PMC article.
References
-
- Motoya T, Nagasawa K, Matsushima Y, Nagata N, Ryo A, Sekizuka T, Yamashita A, Kuroda M, Morita Y, Suzuki Y, Sasaki N, Katayama K, Kimura H. Molecular evolution of the VP1 gene in human norovirus GII.4 variants in 1974–2015. Front Microbiol. 2017;8:2399. doi: 10.3389/fmicb.2017.02399. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

