The Cbk1-Ace2 axis guides Candida albicans from yeast to hyphae and back again
- PMID: 33433733
- PMCID: PMC8139900
- DOI: 10.1007/s00294-020-01152-1
The Cbk1-Ace2 axis guides Candida albicans from yeast to hyphae and back again
Abstract
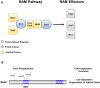
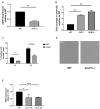
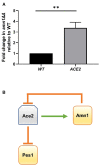
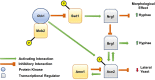
Since its description in S. cerevisiae, the Regulation of Ace2 and Morphogenesis (RAM) pathway has been studied for nearly 20 years in multiple model and pathogenic fungi. In pathogenic fungi, the RAM pathway carries out many functions through mechanisms that remain to be defined in detail. Recently, we reported that Cbk1-mediated phosphorylation of the transcription factor Ace2 functions to repress the hyphae-to-yeast transition in Candida albicans. This transition is understudied relative to the yeast-to-hyphae transition. Subapical hyphal cell compartments are arrested in G1 until the point at which lateral yeast emerge. Here, we discuss this model and report new data indicating that a second G1 associated protein, the mitotic exit regulator Amn1. In S. cerevisiae diploid cells, Amn1 negatively regulates Ace2 at both the gene expression level through a negative feedback loop and at the protein level by targeting Ace2 for degradation. In C. albicans, Amn1 and Ace2 also form a feedback loop at the level of gene expression. Deletion of AMN1 decreases lateral yeast formation relative to wild type in maturing hyphae and is associated with decreased expression of PES1, a positive regulator of lateral yeast formation. These data indicate that the regulation of mitotic exit plays a role in determining the timing of lateral yeast emergence from hyphae in C. albicans. We also propose an integrated model for the interplay between the Cbk1-Ace2 axis and other hyphal stage regulators during the process of filamentation and transition back to yeast.
Keywords: Candida albicans; G1; Hyphae; Lateral yeast; Mitotic exit; RAM network.
Conflict of interest statement
The authors declare no competing interests. All new data are reported in the text of the manuscript. Strains are available by contacting the corresponding author.
Figures

References
-
- Bharucha N, Chabrier-Rosello Y, Xu T, Johnson C, Sobczynski S, Song Q, Dobry CJ, Eckwahl MJ, Anderson CP, Benjamin AJ, Kumar A, Krysan DJ. A large-scale complex haploinsufficiency-based genetic interaction screen in Candida albicans: analysis of the RAM network during morphogenesis. PLoS Genet. 2011;7:e1002058. doi: 10.1371/journal.pgen.1002058. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

