Genomic instability-derived plasma extracellular vesicle-microRNA signature as a minimally invasive predictor of risk and unfavorable prognosis in breast cancer
- PMID: 33436002
- PMCID: PMC7802300
- DOI: 10.1186/s12951-020-00767-3
Genomic instability-derived plasma extracellular vesicle-microRNA signature as a minimally invasive predictor of risk and unfavorable prognosis in breast cancer
Abstract
Background: Breast cancer (BC) is the most frequently diagnosed cancer and the leading cause of cancer-associated deaths in women. Recent studies have indicated that microRNA (miRNA) regulation in genomic instability (GI) is associated with disease risk and clinical outcome. Herein, we aimed to identify the GI-derived miRNA signature in extracellular vesicles (EVs) as a minimally invasive biomarker for early diagnosis and prognostic risk stratification.
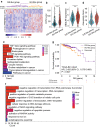
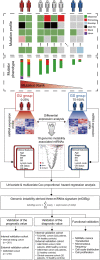
Experimental design: Integrative analysis of miRNA expression and somatic mutation profiles was performed to identify GI-associated miRNAs. Then, we constructed a discovery and validation study with multicenter prospective cohorts. The GI-derived miRNA signature (miGISig) was developed in the TCGA discovery cohort (n = 261), and was subsequently independently validated in internal TCGA validation (n = 261) and GSE22220 (n = 210) cohorts for prognosis prediction, and in GSE73002 (n = 3966), GSE41922 (n = 54), and in-house clinical exosome (n = 30) cohorts for diagnostic performance.
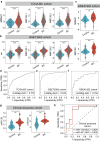
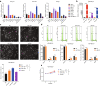
Results: We identified a GI-derived three miRNA signature (MIR421, MIR128-1 and MIR128-2) in the serum extracellular vesicles of BC patients, which was significantly associated with poor prognosis in all the cohorts tested and remained as an independent prognostic factor using multivariate analyses. When integrated with the clinical characteristics, the composite miRNA-clinical prognostic indicator showed improved prognostic performance. The miGISig also showed high accuracy in differentiating BC from healthy controls with the area under the receiver operating characteristics curve (ROC) with 0.915, 0.794 and 0.772 in GSE73002, GSE41922 and TCGA cohorts, respectively. Furthermore, circulating EVs from BC patients in the in-house cohort harbored elevated levels of miGISig, with effective diagnostic accuracy.
Conclusions: We report a novel GI-derived three miRNA signature in EVs, as an excellent minimally invasive biomarker for the early diagnosis and unfavorable prognosis in BC.
Keywords: Breast cancer; Exosomes; Extracellular vesicle; Genomic instability; microRNA.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


References
-
- Bidard FC, Hajage D, Bachelot T, Delaloge S, Brain E, Campone M, et al. Assessment of circulating tumor cells and serum markers for progression-free survival prediction in metastatic breast cancer: a prospective observational study. Breast Cancer Res. 2012;14(1):R29. doi: 10.1186/bcr3114. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

