CTCF and EGR1 suppress breast cancer cell migration through transcriptional control of Nm23-H1
- PMID: 33436746
- PMCID: PMC7804126
- DOI: 10.1038/s41598-020-79869-9
CTCF and EGR1 suppress breast cancer cell migration through transcriptional control of Nm23-H1
Abstract
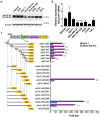
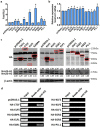
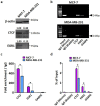
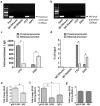
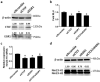
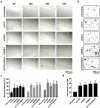
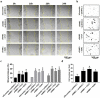
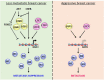
Tumor metastasis remains an obstacle in cancer treatment and is responsible for most cancer-related deaths. Nm23-H1 is one of the first metastasis suppressor proteins discovered with the ability to inhibit metastasis of many cancers including breast, colon, and liver cancer. Although loss of Nm23-H1 is observed in aggressive cancers and correlated with metastatic potential, little is known regarding the mechanisms that regulate its cellular level. Here, we examined the mechanisms that control Nm23-H1 expression in breast cancer cells. Initial studies in aggressive MDA-MB-231 cells (expressing low Nm23-H1) and less invasive MCF-7 cells (expressing high Nm23-H1) revealed that mRNA levels correlated with protein expression, suggesting that transcriptional mechanisms may control Nm23-H1 expression. Truncational analysis of the Nm23-H1 promoter revealed a proximal and minimal promoter that harbor putative binding sites for transcription factors including CTCF and EGR1. CTCF and EGR1 induced Nm23-H1 expression and reduced cell migration of MDA-MB-231 cells. Moreover, CTCF and EGR1 were recruited to the Nm23-H1 promoter in MCF-7 cells and their expression correlated with Nm23-H1 levels. This study indicates that loss of Nm23-H1 in aggressive breast cancer is apparently caused by downregulation of CTCF and EGR1, which potentially drive Nm23-H1 expression to promote a less invasive phenotype.
Conflict of interest statement
The authors declare no competing interests.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

