WLS-Wnt signaling promotes neuroendocrine prostate cancer
- PMID: 33437943
- PMCID: PMC7788232
- DOI: 10.1016/j.isci.2020.101970
WLS-Wnt signaling promotes neuroendocrine prostate cancer
Abstract
Neuroendocrine prostate cancer (NEPC) is a lethal prostate cancer subtype arising as a consequence of more potent androgen receptor (AR) targeting in castration-resistant prostate cancer (CRPC). Its molecular pathogenesis remains elusive. Here, we report that the Wnt secretion mediator Wntless (WLS) is a major driver of NEPC and aggressive tumor growth in vitro and in vivo. Mechanistic studies showed that WLS is a transcriptional target suppressed by AR that activates the ROR2/PKCδ/ERK signaling pathway to support the neuroendocrine (NE) traits and proliferative capacity of NEPC cells. Analysis of clinical samples and datasets revealed that WLS was highly expressed in CRPC and NEPC tumors. Finally, treatment with the Wnt secretion inhibitor LGK974 restricted NE prostate tumor xenograft growth in mice. These findings collectively characterize the contribution of WLS to NEPC pathogenesis and suggest that WLS is a potential therapeutic target in NEPC.
Keywords: Biological Sciences; Cancer; Cell Biology; Transcriptomics.
© 2020 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



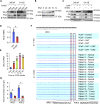




Similar articles
-
The Master Neural Transcription Factor BRN2 Is an Androgen Receptor-Suppressed Driver of Neuroendocrine Differentiation in Prostate Cancer.Cancer Discov. 2017 Jan;7(1):54-71. doi: 10.1158/2159-8290.CD-15-1263. Epub 2016 Oct 26. Cancer Discov. 2017. PMID: 27784708
-
Wntless promotes cellular viability and resistance to enzalutamide in castration-resistant prostate cancer cells.Am J Clin Exp Urol. 2019 Aug 15;7(4):203-214. eCollection 2019. Am J Clin Exp Urol. 2019. PMID: 31511827 Free PMC article.
-
Molecular model for neuroendocrine prostate cancer progression.BJU Int. 2018 Oct;122(4):560-570. doi: 10.1111/bju.14207. Epub 2018 Apr 24. BJU Int. 2018. PMID: 29569310 Review.
-
The central role of Sphingosine kinase 1 in the development of neuroendocrine prostate cancer (NEPC): A new targeted therapy of NEPC.Clin Transl Med. 2022 Feb;12(2):e695. doi: 10.1002/ctm2.695. Clin Transl Med. 2022. PMID: 35184376 Free PMC article.
-
Preclinical Models of Neuroendocrine Prostate Cancer.Curr Protoc. 2023 May;3(5):e742. doi: 10.1002/cpz1.742. Curr Protoc. 2023. PMID: 37166213 Review.
Cited by
-
Aggressive variants of prostate cancer: underlying mechanisms of neuroendocrine transdifferentiation.J Exp Clin Cancer Res. 2022 Feb 2;41(1):46. doi: 10.1186/s13046-022-02255-y. J Exp Clin Cancer Res. 2022. PMID: 35109899 Free PMC article. Review.
-
Monoamine oxidase A: An emerging therapeutic target in prostate cancer.Front Oncol. 2023 Feb 13;13:1137050. doi: 10.3389/fonc.2023.1137050. eCollection 2023. Front Oncol. 2023. PMID: 36860320 Free PMC article. Review.
-
Blocking the Wnt/β‑catenin signaling pathway to treat colorectal cancer: Strategies to improve current therapies (Review).Int J Oncol. 2023 Feb;62(2):24. doi: 10.3892/ijo.2022.5472. Epub 2022 Dec 29. Int J Oncol. 2023. PMID: 36579676 Free PMC article. Review.
-
A novel role for Neurog2 in MYCN driven neuroendocrine plasticity of prostate cancer.Oncogene. 2025 Aug;44(29):2460-2473. doi: 10.1038/s41388-025-03413-0. Epub 2025 Apr 29. Oncogene. 2025. PMID: 40301542 Free PMC article.
-
Neuropilin-2 promotes lineage plasticity and progression to neuroendocrine prostate cancer.Oncogene. 2022 Sep;41(37):4307-4317. doi: 10.1038/s41388-022-02437-0. Epub 2022 Aug 19. Oncogene. 2022. PMID: 35986103 Free PMC article.
References
-
- Akamatsu S., Wyatt A.W., Lin D., Lysakowski S., Zhang F., Kim S., Tse C., Wang K., Mo F., Haegert A. The placental gene PEG10 promotes progression of neuroendocrine prostate cancer. Cell Rep. 2015;12:922–936. - PubMed
-
- Banziger C., Soldini D., Schutt C., Zipperlen P., Hausmann G., Basler K. Wntless, a conserved membrane protein dedicated to the secretion of Wnt proteins from signaling cells. Cell. 2006;125:509–522. - PubMed
-
- Bartscherer K., Pelte N., Ingelfinger D., Boutros M. Secretion of Wnt ligands requires Evi, a conserved transmembrane protein. Cell. 2006;125:523–533. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

