Sex-specific differences in peripheral blood leukocyte transcriptional response to LPS are enriched for HLA region and X chromosome genes
- PMID: 33441806
- PMCID: PMC7806814
- DOI: 10.1038/s41598-020-80145-z
Sex-specific differences in peripheral blood leukocyte transcriptional response to LPS are enriched for HLA region and X chromosome genes
Abstract
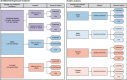
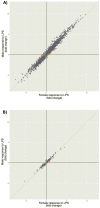
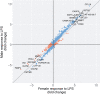
Sex-specific differences in prevalence are well documented for many common, complex diseases, especially for immune-mediated diseases, yet the precise mechanisms through which factors associated with biological sex exert their effects throughout life are not well understood. We interrogated sex-specific transcriptional responses of peripheral blood leukocytes (PBLs) to innate immune stimulation by lipopolysaccharide (LPS) in 46 male and 66 female members of the Hutterite community, who practice a communal lifestyle. We identified 1217 autosomal and 54 X-linked genes with sex-specific responses to LPS, as well as 71 autosomal and one X-linked sex-specific expression quantitative trait loci (eQTLs). Despite a similar proportion of the 15 HLA genes responding to LPS compared to all expressed autosomal genes, there was a significant over-representation of genes with sex by treatment interactions among HLA genes. We also observed an enrichment of sex-specific differentially expressed genes in response to LPS for X-linked genes compared to autosomal genes, suggesting that HLA and X-linked genes may disproportionately contribute to sex disparities in risk for immune-mediated diseases.
Conflict of interest statement
The authors declare no competing interests.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

