DNA Methylation Patterns in T Cells of Naïve and Influenza A Virus-Infected Mice Developmentally Exposed to an Aryl Hydrocarbon Receptor Ligand
- PMID: 33449811
- PMCID: PMC7810290
- DOI: 10.1289/EHP7699
DNA Methylation Patterns in T Cells of Naïve and Influenza A Virus-Infected Mice Developmentally Exposed to an Aryl Hydrocarbon Receptor Ligand
Abstract
Background: Early life environmental exposures can have lasting effects on the function of the immune system and contribute to disease later in life. Epidemiological studies have linked early life exposure to xenobiotics that bind the aryl hydrocarbon receptor (AhR) with dysregulated immune responses later in life. Among the immune cells influenced by developmental activation of the AhR are T cells. Yet, the underlying affected cellular pathways via which activating the AhR early in life causes the responses of T cells to remain affected into adulthood remain unclear.
Objective: Our goal was to identify cellular mechanisms that drive impaired T-cell responses later in life following maternal exposure to an exogenous AhR ligand.
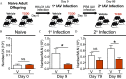
Methods: C57BL/6 mice were vertically exposed to the prototype AhR ligand, 2,3,7,8-tetrachlorodibenzo--dioxin (TCDD), throughout gestation and early postnatal life. The transcriptome and DNA methylation patterns were evaluated in T cells isolated from naïve and influenza A virus (IAV)-infected adult mice that were developmentally exposed to TCDD or vehicle control. We then assessed the influence of DNA methylation-altering drug therapies on the response of T cells from developmentally exposed mice to infection.
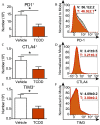
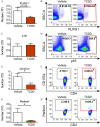
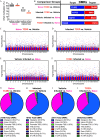
Results: Gene and protein expression showed that developmental AhR activation reduced T-cell expansion and effector functions during IAV infection later in life. Furthermore, whole-genome bisulfite sequencing analyses revealed that developmental AhR activation durably programed DNA methylation patterns across the T-cell genome. Treatment of developmentally exposed offspring with DNA methylation-altering drugs alleviated some, but not all, of the impaired T-cell responses.
Discussion: Taken together, these results indicate that skewed DNA methylation is one of the mechanisms by which early life exposures can durably change the function of T cells in mice. Furthermore, treatment with DNA methylation-altering drugs after the exposure restored some aspects of T-cell functional responsiveness. https://doi.org/10.1289/EHP7699.
Figures

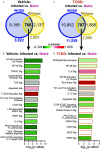
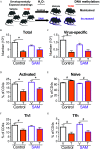
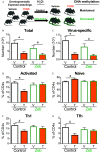
Comment in
-
Developmental Origins of Delayed Adult Immune Response: The AhR Connection.Environ Health Perspect. 2021 May;129(5):54001. doi: 10.1289/EHP9275. Epub 2021 May 12. Environ Health Perspect. 2021. PMID: 33979198 Free PMC article.
Similar articles
-
Effects of developmental activation of the AhR on CD4+ T-cell responses to influenza virus infection in adult mice.Environ Health Perspect. 2014 Nov;122(11):1201-8. doi: 10.1289/ehp.1408110. Epub 2014 Jul 22. Environ Health Perspect. 2014. PMID: 25051576 Free PMC article.
-
Activation of the aryl hydrocarbon receptor during development enhances the pulmonary CD4+ T-cell response to viral infection.Am J Physiol Lung Cell Mol Physiol. 2015 Aug 1;309(3):L305-13. doi: 10.1152/ajplung.00135.2015. Epub 2015 Jun 12. Am J Physiol Lung Cell Mol Physiol. 2015. PMID: 26071552 Free PMC article.
-
Environmental cues received during development shape dendritic cell responses later in life.PLoS One. 2018 Nov 9;13(11):e0207007. doi: 10.1371/journal.pone.0207007. eCollection 2018. PLoS One. 2018. PMID: 30412605 Free PMC article.
-
The aryl hydrocarbon receptor is a modulator of anti-viral immunity.Biochem Pharmacol. 2009 Feb 15;77(4):642-53. doi: 10.1016/j.bcp.2008.10.031. Epub 2008 Nov 5. Biochem Pharmacol. 2009. PMID: 19027719 Free PMC article. Review.
-
Aryl Hydrocarbon Receptor-Dependent Pathways in Immune Regulation.Am J Transplant. 2016 Aug;16(8):2270-6. doi: 10.1111/ajt.13716. Epub 2016 Mar 10. Am J Transplant. 2016. PMID: 26751261 Review.
Cited by
-
Postnatal administration of S-adenosylmethionine restores developmental AHR activation-induced deficits in CD8+ T cell function during influenza A virus infection.Toxicol Sci. 2023 Feb 27;192(2):233-46. doi: 10.1093/toxsci/kfad019. Online ahead of print. Toxicol Sci. 2023. PMID: 36847456 Free PMC article.
-
Developmental Origins of Delayed Adult Immune Response: The AhR Connection.Environ Health Perspect. 2021 May;129(5):54001. doi: 10.1289/EHP9275. Epub 2021 May 12. Environ Health Perspect. 2021. PMID: 33979198 Free PMC article.
-
Consensus on the Key Characteristics of Immunotoxic Agents as a Basis for Hazard Identification.Environ Health Perspect. 2022 Oct;130(10):105001. doi: 10.1289/EHP10800. Epub 2022 Oct 6. Environ Health Perspect. 2022. PMID: 36201310 Free PMC article.
-
Cutaneous Effects of In Utero and Lactational Exposure of C57BL/6J Mice to 2,3,7,8-Tetrachlorodibenzo-p-dioxin.Toxics. 2021 Aug 20;9(8):192. doi: 10.3390/toxics9080192. Toxics. 2021. PMID: 34437510 Free PMC article.
-
Role of aryl hydrocarbon receptors in infection and inflammation.Front Immunol. 2024 Apr 12;15:1367734. doi: 10.3389/fimmu.2024.1367734. eCollection 2024. Front Immunol. 2024. PMID: 38680494 Free PMC article. Review.
References
-
- Barrett T, Inglis SC. 1985. Growth, purification and titration of influenza viruses. In Virology: A Practical Approach. Mahy BWJ, ed. Washington, DC: IRL Press, 119–150.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

