Polymerase γ efficiently replicates through many natural template barriers but stalls at the HSP1 quadruplex
- PMID: 33454015
- PMCID: PMC7762954
- DOI: 10.1074/jbc.RA120.015390
Polymerase γ efficiently replicates through many natural template barriers but stalls at the HSP1 quadruplex
Abstract
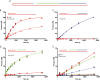
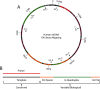
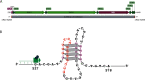
Faithful replication of the mitochondrial genome is carried out by a set of key nuclear-encoded proteins. DNA polymerase γ is a core component of the mtDNA replisome and the only replicative DNA polymerase localized to mitochondria. The asynchronous mechanism of mtDNA replication predicts that the replication machinery encounters dsDNA and unique physical barriers such as structured genes, G-quadruplexes, and other obstacles. In vitro experiments here provide evidence that the polymerase γ heterotrimer is well-adapted to efficiently synthesize DNA, despite the presence of many naturally occurring roadblocks. However, we identified a specific G-quadruplex-forming sequence at the heavy-strand promoter (HSP1) that has the potential to cause significant stalling of mtDNA replication. Furthermore, this structured region of DNA corresponds to the break site for a large (3,895 bp) deletion observed in mitochondrial disease patients. The presence of this deletion in humans correlates with UV exposure, and we have found that efficiency of polymerase γ DNA synthesis is reduced after this quadruplex is exposed to UV in vitro.
Keywords: DNA polymerase γ; DNA replication; DNA structure; G-quadruplex; heavy-strand promoter; mitochondrial DNA (mtDNA); mitochondrial DNA damage; mtDNA deletion.
Copyright © 2020.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health
Figures




Similar articles
-
DNA sequences proximal to human mitochondrial DNA deletion breakpoints prevalent in human disease form G-quadruplexes, a class of DNA structures inefficiently unwound by the mitochondrial replicative Twinkle helicase.J Biol Chem. 2014 Oct 24;289(43):29975-93. doi: 10.1074/jbc.M114.567073. Epub 2014 Sep 5. J Biol Chem. 2014. PMID: 25193669 Free PMC article.
-
Consequences of compromised mitochondrial genome integrity.DNA Repair (Amst). 2020 Sep;93:102916. doi: 10.1016/j.dnarep.2020.102916. DNA Repair (Amst). 2020. PMID: 33087282 Free PMC article. Review.
-
Oxidative DNA damage stalls the human mitochondrial replisome.Sci Rep. 2016 Jul 1;6:28942. doi: 10.1038/srep28942. Sci Rep. 2016. PMID: 27364318 Free PMC article.
-
In vivo occupancy of mitochondrial single-stranded DNA binding protein supports the strand displacement mode of DNA replication.PLoS Genet. 2014 Dec 4;10(12):e1004832. doi: 10.1371/journal.pgen.1004832. eCollection 2014 Dec. PLoS Genet. 2014. PMID: 25474639 Free PMC article.
-
Human mitochondrial DNA replication machinery and disease.Curr Opin Genet Dev. 2016 Jun;38:52-62. doi: 10.1016/j.gde.2016.03.005. Epub 2016 Apr 9. Curr Opin Genet Dev. 2016. PMID: 27065468 Free PMC article. Review.
Cited by
-
Mechanism of strand displacement DNA synthesis by the coordinated activities of human mitochondrial DNA polymerase and SSB.Nucleic Acids Res. 2023 Feb 28;51(4):1750-1765. doi: 10.1093/nar/gkad037. Nucleic Acids Res. 2023. PMID: 36744436 Free PMC article.
-
Interactions of Mitochondrial Transcription Factor A with DNA Damage: Mechanistic Insights and Functional Implications.Genes (Basel). 2021 Aug 15;12(8):1246. doi: 10.3390/genes12081246. Genes (Basel). 2021. PMID: 34440420 Free PMC article. Review.
-
DNA repair pathways in the mitochondria.DNA Repair (Amst). 2025 Feb;146:103814. doi: 10.1016/j.dnarep.2025.103814. Epub 2025 Feb 1. DNA Repair (Amst). 2025. PMID: 39914164 Review.
-
Unravelling How Single-Stranded DNA Binding Protein Coordinates DNA Metabolism Using Single-Molecule Approaches.Int J Mol Sci. 2023 Feb 1;24(3):2806. doi: 10.3390/ijms24032806. Int J Mol Sci. 2023. PMID: 36769124 Free PMC article. Review.
-
G-quadruplex propensity in H. neanderthalensis, H. sapiens and Denisovans mitochondrial genomes.NAR Genom Bioinform. 2024 May 30;6(2):lqae060. doi: 10.1093/nargab/lqae060. eCollection 2024 Jun. NAR Genom Bioinform. 2024. PMID: 38817800 Free PMC article.
References
-
- Anderson S., Bankier A. T., Barrell B. G., de Bruijn M. H., Coulson A. R., Drouin J., Eperon I. C., Nierlich D. P., Roe B. A., Sanger F., Schreier P. H., Smith A. J., Staden R., and Young I. G. (1981) Sequence and organization of the human mitochondrial genome. Nature 290, 457–465 10.1038/290457a0 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

