Establishment and application of information resource of mutant mice in RIKEN BioResource Research Center
- PMID: 33455583
- PMCID: PMC7811887
- DOI: 10.1186/s42826-020-00068-8
Establishment and application of information resource of mutant mice in RIKEN BioResource Research Center
Abstract
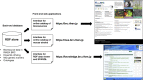
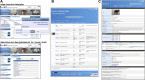
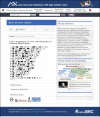
Online databases are crucial infrastructures to facilitate the wide effective and efficient use of mouse mutant resources in life sciences. The number and types of mouse resources have been rapidly growing due to the development of genetic modification technology with associated information of genomic sequence and phenotypes. Therefore, data integration technologies to improve the findability, accessibility, interoperability, and reusability of mouse strain data becomes essential for mouse strain repositories. In 2020, the RIKEN BioResource Research Center released an integrated database of bioresources including, experimental mouse strains, Arabidopsis thaliana as a laboratory plant, cell lines, microorganisms, and genetic materials using Resource Description Framework-related technologies. The integrated database shows multiple advanced features for the dissemination of bioresource information. The current version of our online catalog of mouse strains which functions as a part of the integrated database of bioresources is available from search bars on the page of the Center ( https://brc.riken.jp ) and the Experimental Animal Division ( https://mus.brc.riken.jp/ ) websites. The BioResource Research Center also released a genomic variation database of mouse strains established in Japan and Western Europe, MoG+ ( https://molossinus.brc.riken.jp/mogplus/ ), and a database for phenotype-phenotype associations across the mouse phenome using data from the International Mouse Phenotyping Platform. In this review, we describe features of current version of databases related to mouse strain resources in RIKEN BioResource Research Center and discuss future views.
Keywords: Bioresource; Database; Mouse mutation information resource; Ontology; Semantic web; data integration.
Conflict of interest statement
The authors have no competition of interest directly relevant to the content of this article.
Figures


Similar articles
-
Federated SPARQL query performance evaluation for exploring disease model mouse: combining gene expression, orthology, and disease knowledge graphs.BMC Med Inform Decis Mak. 2025 May 16;25(Suppl 1):189. doi: 10.1186/s12911-025-03013-8. BMC Med Inform Decis Mak. 2025. PMID: 40380154 Free PMC article.
-
Mouse resources at the RIKEN BioResource Research Center and the National BioResource Project core facility in Japan.Mamm Genome. 2022 Mar;33(1):181-191. doi: 10.1007/s00335-021-09916-x. Epub 2021 Sep 16. Mamm Genome. 2022. PMID: 34532769 Free PMC article. Review.
-
The mouse resources at the RIKEN BioResource center.Exp Anim. 2009 Apr;58(2):85-96. doi: 10.1538/expanim.58.85. Exp Anim. 2009. PMID: 19448331 Review.
-
Genetic materials at the gene engineering division, RIKEN BioResource Center.Exp Anim. 2010;59(2):115-24. doi: 10.1538/expanim.59.115. Exp Anim. 2010. PMID: 20484845 Review.
-
RARGE: a large-scale database of RIKEN Arabidopsis resources ranging from transcriptome to phenome.Nucleic Acids Res. 2005 Jan 1;33(Database issue):D647-50. doi: 10.1093/nar/gki014. Nucleic Acids Res. 2005. PMID: 15608280 Free PMC article.
Cited by
-
Principles for establishment of the stem cell bank and its applications on management of sports injuries.Stem Cell Res Ther. 2021 May 29;12(1):307. doi: 10.1186/s13287-021-02360-3. Stem Cell Res Ther. 2021. PMID: 34051865 Free PMC article. Review.
-
Federated SPARQL query performance evaluation for exploring disease model mouse: combining gene expression, orthology, and disease knowledge graphs.BMC Med Inform Decis Mak. 2025 May 16;25(Suppl 1):189. doi: 10.1186/s12911-025-03013-8. BMC Med Inform Decis Mak. 2025. PMID: 40380154 Free PMC article.
-
Asian Mouse Mutagenesis Resource Association (AMMRA): mouse genetics and laboratory animal resources in the Asia Pacific.Mamm Genome. 2022 Mar;33(1):192-202. doi: 10.1007/s00335-021-09912-1. Epub 2021 Sep 5. Mamm Genome. 2022. PMID: 34482437 Free PMC article. Review.
-
Mouse resources at the RIKEN BioResource Research Center and the National BioResource Project core facility in Japan.Mamm Genome. 2022 Mar;33(1):181-191. doi: 10.1007/s00335-021-09916-x. Epub 2021 Sep 16. Mamm Genome. 2022. PMID: 34532769 Free PMC article. Review.
-
MoG+: a database of genomic variations across three mouse subspecies for biomedical research.Mamm Genome. 2022 Mar;33(1):31-43. doi: 10.1007/s00335-021-09933-w. Epub 2021 Nov 15. Mamm Genome. 2022. PMID: 34782917 Free PMC article.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

