Molecular Dynamics Simulation of Atomic Interactions in the Vancomycin Binding Site
- PMID: 33458529
- PMCID: PMC7808135
- DOI: 10.1021/acsomega.0c05353
Molecular Dynamics Simulation of Atomic Interactions in the Vancomycin Binding Site
Abstract
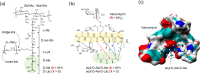
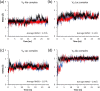
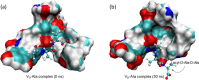
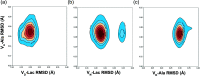
Vancomycin is a glycopeptide antibiotic produced by Amycolaptopsis orientalis used to treat serious infections by Gram-positive pathogens including methicillin-resistant Staphylococcus aureus. Vancomycin inhibits cell wall biosynthesis by targeting lipid II, which is the membrane-bound peptidoglycan precursor. The heptapeptide aglycon structure of vancomycin binds to the d-Ala-d-Ala of the pentapeptide stem structure in lipid II. The third residue of vancomycin aglycon is asparagine, which is not directly involved in the dipeptide binding. Nonetheless, asparagine plays a crucial role in substrate recognition, as the vancomycin analogue with asparagine substituted by aspartic acid (VD) shows a reduction in antibacterial activities. To characterize the function of asparagine, binding of vancomycin and its aspartic-acid-substituted analogue VD to l-Lys-d-Ala-d-Ala and l-Lys-d-Ala-d-Lac was investigated using molecular dynamic simulations. Binding interactions were analyzed using root-mean-square deviation (RMSD), two-dimensional (2D) contour plots, hydrogen bond analysis, and free energy calculations of the complexes. The analysis shows that the aspartate substitution introduced a negative charge to the binding cleft of VD, which altered the aglycon conformation that minimized the repulsive lone pair interaction in the binding of a depsipeptide. Our findings provide new insight for the development of novel glycopeptide antibiotics against the emerging vancomycin-resistant pathogens by chemical modification at the third residue in vancomycin to improve its binding affinity to the d-Ala-d-Lac-terminated peptidoglycan in lipid II found in vancomycin-resistant enterococci and vancomycin-resistant S. aureus.
© 2020 The Authors. Published by American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
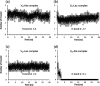
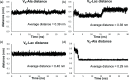
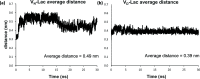
References
-
- McCormick M. H.; McGuire J. M.; Pittenger G. E.; Pittenger R. C.; Stark W. M. Vancomycin, a new antibiotic. I. Chemical and biologic properties. Antibiot. Annu. 1955, 3, 606–611. - PubMed
-
- Nagarajan R.Glycopeptide Antibiotics; Marcel Dekker, New York, 1994; Vol. 63.
-
- Nicolaou K. C.; Hughes R.; Cho S. Y.; Winssinger N.; Labischinski H.; Endermann R. Synthesis and biological evaluation of vancomycin dimers with potent activity against vancomycin-resistant bacteria: target-accelerated combinatorial synthesis. Chemistry 2001, 7, 3824–3843. 10.1002/1521-3765(20010903)7:17<3824::AID-CHEM3824>3.0.CO;2-1. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

