Identification of host factors binding to dengue and Zika virus subgenomic RNA by efficient yeast three-hybrid screens of the human ORFeome
- PMID: 33459164
- PMCID: PMC8086697
- DOI: 10.1080/15476286.2020.1868754
Identification of host factors binding to dengue and Zika virus subgenomic RNA by efficient yeast three-hybrid screens of the human ORFeome
Abstract
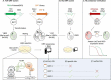
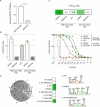
Flaviviruses such as the dengue (DENV) and the Zika virus (ZIKV) are important human pathogens causing around 100 million symptomatic infections each year. During infection, small subgenomic flavivirus RNAs (sfRNAs) are formed inside the infected host cell as a result of incomplete degradation of the viral RNA genome by cellular exoribonuclease XRN1. Although the full extent of sfRNA functions is to be revealed, these non-coding RNAs are key virulence factors and their detrimental effects on multiple cellular processes seem to consistently involve molecular interactions with RNA-binding proteins (RBPs). Discovery of such sfRNA-binding host-factors has followed established biochemical pull-down approaches skewed towards highly abundant proteins hampering proteome-wide coverage. Yeast three-hybrid (Y3H) systems represent an attractive alternative approach. To facilitate proteome-wide screens for RBP, we revisited and improved existing RNA-Y3H methodology by (1) implementing full-length ORF libraries in combination with (2) efficient yeast mating to increase screening depth and sensitivity, and (3) stringent negative controls to eliminate over-representation of non-specific RNA-binders. These improvements were validated employing the well-characterized interaction between DDX6 (DEAD-box helicase 6) and sfRNA of DENV as paradigm. Our advanced Y3H system was used to screen for human proteins binding to DENV and ZIKV sfRNA, resulting in a list of 69 putative sfRNA-binders, including several previously reported as well as numerous novel RBP host factors. Our methodology requiring no sophisticated infrastructure or analytic pipeline may be employed for the discovery of meaningful RNA-protein interactions at large scale in other fields.
Keywords: ORF library; RNA-binding proteins; Subgenomic flavivirus RNA; Zika virus; dengue virus; host factors; yeast three-hybrid.
Conflict of interest statement
No potential conflicts of interest were disclosed.
Figures


Similar articles
-
The subgenomic flaviviral RNA suppresses RNA interference through competing with siRNAs for binding RISC components.J Virol. 2024 Feb 20;98(2):e0195423. doi: 10.1128/jvi.01954-23. Epub 2024 Jan 30. J Virol. 2024. PMID: 38289102 Free PMC article.
-
Subgenomic Flaviviral RNAs of Dengue Viruses.Viruses. 2023 Nov 24;15(12):2306. doi: 10.3390/v15122306. Viruses. 2023. PMID: 38140548 Free PMC article. Review.
-
Zika Virus Subgenomic Flavivirus RNA Generation Requires Cooperativity between Duplicated RNA Structures That Are Essential for Productive Infection in Human Cells.J Virol. 2020 Aug 31;94(18):e00343-20. doi: 10.1128/JVI.00343-20. Print 2020 Aug 31. J Virol. 2020. PMID: 32581095 Free PMC article.
-
Zika virus noncoding sfRNAs sequester multiple host-derived RNA-binding proteins and modulate mRNA decay and splicing during infection.J Biol Chem. 2019 Nov 1;294(44):16282-16296. doi: 10.1074/jbc.RA119.009129. Epub 2019 Sep 13. J Biol Chem. 2019. PMID: 31519749 Free PMC article.
-
Functional non-coding RNAs derived from the flavivirus 3' untranslated region.Virus Res. 2015 Aug 3;206:53-61. doi: 10.1016/j.virusres.2015.01.026. Epub 2015 Feb 7. Virus Res. 2015. PMID: 25660582 Review.
Cited by
-
Integrated multi-omics analyses identify key anti-viral host factors and pathways controlling SARS-CoV-2 infection.Res Sq [Preprint]. 2022 Aug 15:rs.3.rs-1910932. doi: 10.21203/rs.3.rs-1910932/v1. Res Sq. 2022. Update in: Nat Commun. 2024 Jan 2;15(1):109. doi: 10.1038/s41467-023-44175-1. PMID: 36032971 Free PMC article. Updated. Preprint.
-
The subgenomic flaviviral RNA suppresses RNA interference through competing with siRNAs for binding RISC components.J Virol. 2024 Feb 20;98(2):e0195423. doi: 10.1128/jvi.01954-23. Epub 2024 Jan 30. J Virol. 2024. PMID: 38289102 Free PMC article.
-
Subgenomic Flaviviral RNAs of Dengue Viruses.Viruses. 2023 Nov 24;15(12):2306. doi: 10.3390/v15122306. Viruses. 2023. PMID: 38140548 Free PMC article. Review.
-
Host cytoskeletal vimentin serves as a structural organizer and an RNA-binding protein regulator to facilitate Zika viral replication.Proc Natl Acad Sci U S A. 2022 Feb 22;119(8):e2113909119. doi: 10.1073/pnas.2113909119. Proc Natl Acad Sci U S A. 2022. PMID: 35193960 Free PMC article.
-
CRISPR Screen Reveals PACT as a Pro-Viral Factor for Dengue Viral Replication.Viruses. 2024 May 3;16(5):725. doi: 10.3390/v16050725. Viruses. 2024. PMID: 38793607 Free PMC article.
References
-
- Riva P, Ratti A, Venturin M.. The long non-coding RNAs in neurodegenerative diseases: novel mechanisms of pathogenesis. Curr Alzheimer Res. 2016;13:1219–1231. - PubMed
-
- Hentze MW, Castello A, Schwarzl T, et al. A brave new world of RNA-binding proteins. Nat Rev Mol Cell Biol. 2018;19:327–341. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
