Molecular detection of Leishmania donovani, Leishmania major, and Trypanosoma species in Sergentomyia squamipleuris sand flies from a visceral leishmaniasis focus in Merti sub-County, eastern Kenya
- PMID: 33461609
- PMCID: PMC7812738
- DOI: 10.1186/s13071-020-04517-0
Molecular detection of Leishmania donovani, Leishmania major, and Trypanosoma species in Sergentomyia squamipleuris sand flies from a visceral leishmaniasis focus in Merti sub-County, eastern Kenya
Abstract
Background: Visceral leishmaniasis (VL) and zoonotic cutaneous leishmaniasis (ZCL) are of public health concern in Merti sub-County, Kenya, but epidemiological data on transmission, vector abundance, distribution, and reservoir hosts remain limited. To better understand the disease and inform control measures to reduce transmission, we investigated the abundance and distribution of sand fly species responsible for Leishmania transmission in the sub-County and their blood-meal hosts.
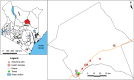
Methods: We conducted an entomological survey in five villages with reported cases of VL in Merti sub-County, Kenya, using CDC miniature light traps and castor oil sticky papers. Sand flies were dissected and identified to the species level using standard taxonomic keys and PCR analysis of the cytochrome c oxidase subunit 1 (cox1) gene. Leishmania parasites were detected and identified by PCR and sequencing of internal transcribed spacer 1 (ITS1) genes. Blood-meal sources of engorged females were identified by high-resolution melting analysis of vertebrate cytochrome b (cyt-b) gene PCR products.
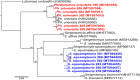
Results: We sampled 526 sand flies consisting of 8 species, Phlebotomus orientalis (1.52%; n = 8), and 7 Sergentomyia spp. Sergentomyia squamipleuris was the most abundant sand fly species (78.71%; n = 414) followed by Sergentomyia clydei (10.46%; n = 55). Leishmania major, Leishmania donovani, and Trypanosoma DNA were detected in S. squamipleuris specimens. Humans were the main sources of sand fly blood meals. However, we also detected mixed blood meals; one S. squamipleuris specimen had fed on both human and mouse (Mus musculus) blood, while two Ph. orientalis specimens fed on human, hyrax (Procavia capensis), and mouse (Mus musculus) blood.
Conclusions: Our findings implicate the potential involvement of S. squamipleuris in the transmission of Leishmania and question the dogma that human leishmaniases in the Old World are exclusively transmitted by sand flies of the Phlebotomus genus. The presence of Trypanosoma spp. may indicate mechanical transmission, whose efficiency should be investigated. Host preference analysis revealed the possibility of zoonotic transmission of leishmaniasis and other pathogens in the sub-County. Leishmania major and L. donovani are known to cause ZCL and VL, respectively. However, the reservoir status of the parasites is not uniform. Further studies are needed to determine the reservoir hosts of Leishmania spp. in the area.
Keywords: Kenya; Leishmania; Leishmaniasis; Phlebotomus; Sand fly; Sergentomyia; Trypanosoma; Visceral leishmaniasis; Zoonotic cutaneous leishmaniasis.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


References
-
- Nzelu CO, Cáceres AG, Arrunátegui-Jiménez MJ, Lañas-Rosas MF, Yañez-Trujillano HH, Luna-Caipo DV, et al. DNA barcoding for identification of sand fly species (Diptera: Psychodidae) from leishmaniasis-endemic areas of Peru. Acta Trop. 2015;145:45–51. doi: 10.1016/j.actatropica.2015.02.003. - DOI - PubMed
-
- Owino BO, Matoke-Muhia D, Alraey Y, Mwangi JM, Ingonga JM, Ngumbi PM, et al. Association of Phlebotomus guggisbergi with Leishmania major and Leishmania tropica in a complex transmission setting for cutaneous leishmaniasis in Gilgil, Nakuru County, Kenya. PLoS Negl Trop Dis. 2019;13(10):1–21. doi: 10.1371/journal.pntd.0007712. - DOI - PMC - PubMed
-
- World Health Organization (WHO). Leishmaniasis 2019. https://www.who.int/en/news-room/fact-sheets/detail/leishmaniasis.
-
- Njau J. Leishmaniasis control in Kenya: current challenges and future strategies. 2010.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

