Freshwater monitoring by nanopore sequencing
- PMID: 33461660
- PMCID: PMC7815314
- DOI: 10.7554/eLife.61504
Freshwater monitoring by nanopore sequencing
Abstract
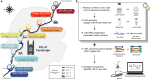
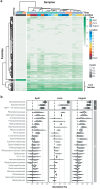
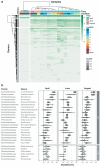
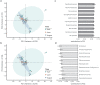
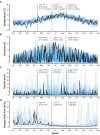
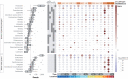
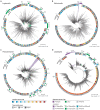
While traditional microbiological freshwater tests focus on the detection of specific bacterial indicator species, including pathogens, direct tracing of all aquatic DNA through metagenomics poses a profound alternative. Yet, in situ metagenomic water surveys face substantial challenges in cost and logistics. Here, we present a simple, fast, cost-effective and remotely accessible freshwater diagnostics workflow centred around the portable nanopore sequencing technology. Using defined compositions and spatiotemporal microbiota from surface water of an example river in Cambridge (UK), we provide optimised experimental and bioinformatics guidelines, including a benchmark with twelve taxonomic classification tools for nanopore sequences. We find that nanopore metagenomics can depict the hydrological core microbiome and fine temporal gradients in line with complementary physicochemical measurements. In a public health context, these data feature relevant sewage signals and pathogen maps at species level resolution. We anticipate that this framework will gather momentum for new environmental monitoring initiatives using portable devices.
Keywords: bacterial monitoring; computational biology; ecology; environmental metagenomics; freshwater ecology; infectious disease; microbiology; nanopore sequencing; portable dna analysis.
Plain language summary
Many water-dwelling bacteria can cause severe diseases such as cholera, typhoid or leptospirosis. One way to prevent outbreaks is to test water sources to find out which species of microbes they contain, and at which levels. Traditionally, this involves taking a water sample, followed by growing a few species of ‘indicator bacteria’ that help to estimate whether the water is safe. An alternative technique, called metagenomics, has been available since the mid-2000s. It consists in reviewing (or ‘sequencing’) the genetic information of most of the bacteria present in the water, which allows scientists to spot harmful species. Both methods, however, require well-equipped laboratories with highly trained staff, making them challenging to use in remote areas. The MinION is a pocket-sized device that – when paired with a laptop or mobile phone – can sequence genetic information ‘on the go’. It has already been harnessed during Ebola, Zika or SARS-CoV-2 epidemics to track the genetic information of viruses in patients and environmental samples. However, it is still difficult to use the MinION and other sequencers to monitor bacteria in water sources, partly because the genetic information of the microbes is highly fragmented during DNA extraction. To address this challenge, Urban, Holzer et al. set out to optimise hardware and software protocols so the MinION could be used to detect bacterial species present in rivers. The tests focussed on the River Cam in Cambridge, UK, a waterway which faces regular public health problems: local rowers and swimmers often contract waterborne infections, sometimes leading to river closures. For six months, Urban, Holzer et al. used the MinION to map out the bacteria present across nine river sites, assessing the diversity of species and the presence of disease-causing microbes in the water. In particular, the results showed that optimising the protocols made it possible to tell the difference between closely related species – an important feature since harmful and inoffensive bacteria can sometimes be genetically close. The data also revealed that the levels of harmful bacteria were highest downstream of urban river sections, near a water treatment plant and river barge moorings. Together, these findings demonstrate that optimising MinION protocols can turn this device into a useful tool to easily monitor water quality. Around the world, climate change, rising urbanisation and the intensification of agriculture all threaten water quality. In fact, access to clean water is one of the United Nations sustainable development goals for 2030. Using the guidelines developed by Urban, Holzer et al., communities could harness the MinION to monitor water quality in remote areas, offering a cost-effective, portable DNA analysis tool to protect populations against deadly diseases.
© 2021, Urban et al.
Conflict of interest statement
LU, AH, JB, MH, PB, MS, DK, SP, DM, ET, SS, MS No competing interests declared
Figures








References
Publication types
MeSH terms
Substances
Grants and funding
- OpenPlant Fund (BBSRC BB/L014130/1)/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- Graduate Student Fellowship (203828/Z/16/A, 203828/Z/16/Z)/WT_/Wellcome Trust/United Kingdom
- WT_/Wellcome Trust/United Kingdom
- Graduate Student Fellowship (102453/Z/13/Z)/WT_/Wellcome Trust/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

