In vivo nanoscale analysis of the dynamic synergistic interaction of Bacillus thuringiensis Cry11Aa and Cyt1Aa toxins in Aedes aegypti
- PMID: 33465145
- PMCID: PMC7846010
- DOI: 10.1371/journal.ppat.1009199
In vivo nanoscale analysis of the dynamic synergistic interaction of Bacillus thuringiensis Cry11Aa and Cyt1Aa toxins in Aedes aegypti
Abstract
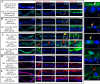
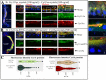
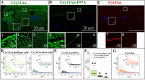
The insecticidal Cry11Aa and Cyt1Aa proteins are produced by Bacillus thuringiensis as crystal inclusions. They work synergistically inducing high toxicity against mosquito larvae. It was proposed that these crystal inclusions are rapidly solubilized and activated in the gut lumen, followed by pore formation in midgut cells killing the larvae. In addition, Cyt1Aa functions as a Cry11Aa binding receptor, inducing Cry11Aa oligomerization and membrane insertion. Here, we used fluorescent labeled crystals, protoxins or activated toxins for in vivo localization at nano-scale resolution. We show that after larvae were fed solubilized proteins, these proteins were not accumulated inside the gut and larvae were not killed. In contrast, if larvae were fed soluble non-toxic mutant proteins, these proteins were found inside the gut bound to gut-microvilli. Only feeding with crystal inclusions resulted in high larval mortality, suggesting that they have a role for an optimal intoxication process. At the macroscopic level, Cry11Aa completely degraded the gastric caeca structure and, in the presence of Cyt1Aa, this effect was observed at lower toxin-concentrations and at shorter periods. The labeled Cry11Aa crystal protein, after midgut processing, binds to the gastric caeca and posterior midgut regions, and also to anterior and medium regions where it is internalized in ordered "net like" structures, leading finally to cell break down. During synergism both Cry11Aa and Cyt1Aa toxins showed a dynamic layered array at the surface of apical microvilli, where Cry11Aa is localized in the lower layer closer to the cell cytoplasm, and Cyt1Aa is layered over Cry11Aa. This array depends on the pore formation activity of Cry11Aa, since the non-toxic mutant Cry11Aa-E97A, which is unable to oligomerize, inverted this array. Internalization of Cry11Aa was also observed during synergism. These data indicate that the mechanism of action of Cry11Aa is more complex than previously anticipated, and may involve additional steps besides pore-formation activity.
Conflict of interest statement
The authors have declared that no competing interest exist in this work.
Figures



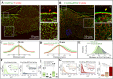
References
-
- Dadd H. Effects of size and concentration of particles on rates of ingestion of latex particulates by mosquito larvae. Annals of the Entomological Society of America. 1971; 64: 687–693.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

