PE_PGRS33, an Important Virulence Factor of Mycobacterium tuberculosis and Potential Target of Host Humoral Immune Response
- PMID: 33467487
- PMCID: PMC7830552
- DOI: 10.3390/cells10010161
PE_PGRS33, an Important Virulence Factor of Mycobacterium tuberculosis and Potential Target of Host Humoral Immune Response
Abstract
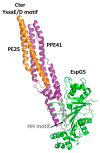
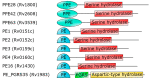
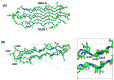
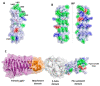
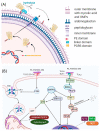
PE_PGRS proteins are surface antigens of Mycobacterium tuberculosis (Mtb) and a few other pathogenic mycobacteria. The PE_PGRS33 protein is among the most studied PE_PGRSs. It is known that the PE domain of PE_PGRS33 is required for the protein translocation through the mycobacterial cell wall, where the PGRS domain remains available for interaction with host receptors. Interaction with Toll like receptor 2 (TLR2) promotes secretion of inflammatory chemokines and cytokines, which are key in the immunopathogenesis of tuberculosis (TB). In this review, we briefly address some key challenges in the development of a TB vaccine and attempt to provide a rationale for the development of new vaccines aimed at fostering a humoral response against Mtb. Using PE_PGRS33 as a model for a surface-exposed antigen, we exploit the availability of current structural data using homology modeling to gather insights on the PGRS domain features. Our study suggests that the PGRS domain of PE_PGRS33 exposes four PGII sandwiches on the outer surface, which, we propose, are directly involved through their loops in the interactions with the host receptors and, as such, are promising targets for a vaccination strategy aimed at inducing a humoral response.
Keywords: infectious disease; protein structure; tuberculosis; vaccine.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures

References
-
- World Health Organization The End TB Strategy. [(accessed on 14 January 2021)]; Available online: https://www.who.int/tb/strategy/en/
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

