Tetracycline Resistance Mediated by tet(M) Has Variable Integrative Conjugative Element Composition in Mycoplasma hominis Strains Isolated in the United Kingdom from 2005 to 2015
- PMID: 33468475
- PMCID: PMC8097440
- DOI: 10.1128/AAC.02513-20
Tetracycline Resistance Mediated by tet(M) Has Variable Integrative Conjugative Element Composition in Mycoplasma hominis Strains Isolated in the United Kingdom from 2005 to 2015
Abstract
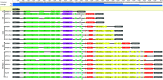
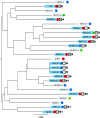
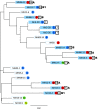
A minimal genome and absent bacterial cell wall render Mycoplasma hominis inherently resistant to most antimicrobials except lincosamides, tetracyclines, and fluoroquinolones. Often dismissed as a commensal (except where linked to preterm birth), it causes septic arthritis in immunodeficient patients and is increasingly associated with transplant failure (particularly lung) accompanying immunosuppression. We examined antimicrobial susceptibility (AST) on strains archived from 2005 to 2015 submitted to the Public Health England reference laboratory and determined the underlying mechanism of resistance by whole-genome sequencing (WGS). Archived M. hominis strains included 32/115 from invasive infection (sepsis, cerebrospinal [CSF], peritoneal, and pleural fluid) over the 10-year period (6.4% of all samples submitted from 2010 to 2015 were positive). No clindamycin resistance was detected, while two strains were resistant to moxifloxacin and levofloxacin (resistance mutations S83L or E87G in gyrA and S81I or E84V in parC). One of these strains and 11 additional strains were tetracycline resistant, mediated by tet(M) carried within an integrative conjugative element (ICE) consistently integrated at the somatic rumA gene; however, the ICEs varied widely in 5 to 19 associated accessory genes. WGS analysis showed that tet(M)-carrying strains were not clonal, refuting previous speculation that the ICE was broken and immobile. We found tet(M)-positive and -negative strains (including the multiresistant 2015 strain) to be equally susceptible to tigecycline and josamycin; however, the British National Formulary does not include guidance for these. Continued M. hominis investigation and AST surveillance (especially immunocompromised patients) is warranted, and the limited number of therapeutics needs to be expanded in the United Kingdom.
Keywords: Mycoplasma; Mycoplasma hominis; United Kingdom; antibiotic resistance; antimicrobial activity; antimicrobial resistance; epidemiology; genome analysis; genomics; integral conjugative element; tetracyclines.
Copyright © 2021 American Society for Microbiology.
Figures



References
-
- Thorsen P, Jensen IP, Jeune B, Ebbesen N, Arpi M, Bremmelgaard A, Møller BR. 1998. Few microorganisms associated with bacterial vaginosis may constitute the pathologic core: a population-based microbiologic study among 3596 pregnant women. Am J Obstet Gynecol 178:580–587. doi:10.1016/s0002-9378(98)70442-9. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

