Genome Sequencing of Sewage Detects Regionally Prevalent SARS-CoV-2 Variants
- PMID: 33468686
- PMCID: PMC7845645
- DOI: 10.1128/mBio.02703-20
Genome Sequencing of Sewage Detects Regionally Prevalent SARS-CoV-2 Variants
Abstract
Viral genome sequencing has guided our understanding of the spread and extent of genetic diversity of SARS-CoV-2 during the COVID-19 pandemic. SARS-CoV-2 viral genomes are usually sequenced from nasopharyngeal swabs of individual patients to track viral spread. Recently, RT-qPCR of municipal wastewater has been used to quantify the abundance of SARS-CoV-2 in several regions globally. However, metatranscriptomic sequencing of wastewater can be used to profile the viral genetic diversity across infected communities. Here, we sequenced RNA directly from sewage collected by municipal utility districts in the San Francisco Bay Area to generate complete and nearly complete SARS-CoV-2 genomes. The major consensus SARS-CoV-2 genotypes detected in the sewage were identical to clinical genomes from the region. Using a pipeline for single nucleotide variant calling in a metagenomic context, we characterized minor SARS-CoV-2 alleles in the wastewater and detected viral genotypes which were also found within clinical genomes throughout California. Observed wastewater variants were more similar to local California patient-derived genotypes than they were to those from other regions within the United States or globally. Additional variants detected in wastewater have only been identified in genomes from patients sampled outside California, indicating that wastewater sequencing can provide evidence for recent introductions of viral lineages before they are detected by local clinical sequencing. These results demonstrate that epidemiological surveillance through wastewater sequencing can aid in tracking exact viral strains in an epidemic context.
Keywords: coronavirus; environmental microbiology; genomics; metagenomics.
Copyright © 2021 Crits-Christoph et al.
Figures


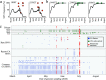
References
-
- Jorden MA, Rudman SL, Villarino E, Hoferka S, Patel MT, Bemis K, Simmons CR, Jespersen M, Iberg Johnson J, Mytty E, Arends KD, Henderson JJ, Mathes RW, Weng CX, Duchin J, Lenahan J, Close N, Bedford T, Boeckh M, Chu HY, Englund JA, Famulare M, Nickerson DA, Rieder MJ, Shendure J, Starita LM, Team CC-19 R, CDC COVID-19 Response Team. 2020. Evidence for Limited Early Spread of COVID-19 Within the United States, January–February 2020. MMWR Morb Mortal Wkly Rep 69:680–684. doi: 10.15585/mmwr.mm6922e1. - DOI - PMC - PubMed
-
- Deng X, Gu W, Federman S, Du Plessis L, Pybus OG, Faria NR, Wang C, Yu G, Bushnell B, Pan C-Y, Guevara H, Sotomayor-Gonzalez A, Zorn K, Gopez A, Servellita V, Hsu E, Miller S, Bedford T, Greninger AL, Roychoudhury P, Starita LM, Famulare M, Chu HY, Shendure J, Jerome KR, Anderson C, Gangavarapu K, Zeller M, Spencer E, Andersen KG, MacCannell D, Paden CR, Li Y, Zhang J, Tong S, Armstrong G, Morrow S, Willis M, Matyas BT, Mase S, Kasirye O, Park M, Masinde G, Chan C, Yu AT, Chai SJ, Villarino E, Bonin B, Wadford DA, Chiu CY, et al. 2020. Genomic surveillance reveals multiple introductions of SARS-CoV-2 into northern California. Science 369:582–587. doi: 10.1126/science.abb9263. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous
