The Canonical Long-Chain Fatty Acid Sensing Machinery Processes Arachidonic Acid To Inhibit Virulence in Enterohemorrhagic Escherichia coli
- PMID: 33468701
- PMCID: PMC7845647
- DOI: 10.1128/mBio.03247-20
The Canonical Long-Chain Fatty Acid Sensing Machinery Processes Arachidonic Acid To Inhibit Virulence in Enterohemorrhagic Escherichia coli
Abstract
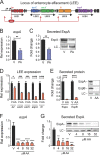
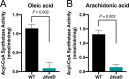
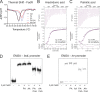
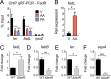
The mammalian gastrointestinal tract is a complex biochemical organ that generates a diverse milieu of host- and microbe-derived metabolites. In this environment, bacterial pathogens sense and respond to specific stimuli, which are integrated into the regulation of their virulence programs. Previously, we identified the transcription factor FadR, a long-chain fatty acid (LCFA) acyl coenzyme A (acyl-CoA) sensor, as a novel virulence regulator in the human foodborne pathogen enterohemorrhagic Escherichia coli (EHEC). Here, we demonstrate that exogenous LCFAs directly inhibit the locus of enterocyte effacement (LEE) pathogenicity island in EHEC through sensing by FadR. Moreover, in addition to LCFAs that are 18 carbons in length or shorter, we introduce host-derived arachidonic acid (C20:4) as an additional LCFA that is recognized by the FadR system in EHEC. We show that arachidonic acid is processed by the acyl-CoA synthetase FadD, which permits binding to FadR and decreases FadR affinity for its target DNA sequences. This interaction enables the transcriptional regulation of FadR-responsive operons by arachidonic acid in EHEC, including the LEE. Finally, we show that arachidonic acid inhibits hallmarks of EHEC disease in a FadR-dependent manner, including EHEC attachment to epithelial cells and the formation of attaching and effacing lesions. Together, our findings delineate a molecular mechanism demonstrating how LCFAs can directly inhibit the virulence of an enteric bacterial pathogen. More broadly, our findings expand the repertoire of ligands sensed by the canonical LFCA sensing machinery in EHEC to include arachidonic acid, an important bioactive lipid that is ubiquitous within host environments.IMPORTANCE Polyunsaturated fatty acids (PUFAs) play important roles in host immunity. Manipulation of lipid content in host tissues through diet or pharmacological interventions is associated with altered severity of various inflammatory diseases. Our work introduces a defined host-pathogen interaction by which arachidonic acid, a host-derived and dietary PUFA, can impact the outcome of enteric infection with the human pathogen enterohemorrhagic Escherichia coli (EHEC). We show that long-chain fatty acids including arachidonic acid act as signaling molecules that directly suppress a key pathogenicity island in EHEC following recognition by the fatty acyl-CoA-responsive transcription factor FadR. Thus, in addition to its established effects on host immunity and its bactericidal activities against other pathogens, we demonstrate that arachidonic acid also acts as a signaling molecule that inhibits virulence in an enteric pathogen.
Keywords: FadR; PUFA; arachidonic acid; enterohemorrhagic E. coli (EHEC); fatty acid; host-pathogen interactions; infection; locus of enterocyte effacement (LEE); omega 6; virulence regulation.
Copyright © 2021 Ellermann et al.
Figures


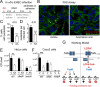
Similar articles
-
Virulence meets metabolism: Cra and KdpE gene regulation in enterohemorrhagic Escherichia coli.mBio. 2012 Oct 16;3(5):e00280-12. doi: 10.1128/mBio.00280-12. mBio. 2012. PMID: 23073764 Free PMC article.
-
Bacterial Adrenergic Sensors Regulate Virulence of Enteric Pathogens in the Gut.mBio. 2016 Jun 7;7(3):e00826-16. doi: 10.1128/mBio.00826-16. mBio. 2016. PMID: 27273829 Free PMC article.
-
Phosphotyrosine-Mediated Regulation of Enterohemorrhagic Escherichia coli Virulence.mBio. 2018 Feb 27;9(1):e00097-18. doi: 10.1128/mBio.00097-18. mBio. 2018. PMID: 29487233 Free PMC article.
-
Key roles of two-component systems in intestinal signal sensing and virulence regulation in enterohemorrhagic Escherichia coli.FEMS Microbiol Rev. 2024 Nov 23;48(6):fuae028. doi: 10.1093/femsre/fuae028. FEMS Microbiol Rev. 2024. PMID: 39537200 Free PMC article. Review.
-
The Tip of the Iceberg: On the Roles of Regulatory Small RNAs in the Virulence of Enterohemorrhagic and Enteropathogenic Escherichia coli.Front Cell Infect Microbiol. 2016 Sep 21;6:105. doi: 10.3389/fcimb.2016.00105. eCollection 2016. Front Cell Infect Microbiol. 2016. PMID: 27709103 Free PMC article. Review.
Cited by
-
Fatty Acid Homeostasis Tunes Flagellar Motility by Activating Phase 2 Flagellin Expression, Contributing to Salmonella Gut Colonization.Infect Immun. 2022 Jul 21;90(7):e0018422. doi: 10.1128/iai.00184-22. Epub 2022 Jun 2. Infect Immun. 2022. PMID: 35652649 Free PMC article.
-
Targeting microbial quorum sensing: the next frontier to hinder bacterial driven gastrointestinal infections.Gut Microbes. 2023 Dec;15(2):2252780. doi: 10.1080/19490976.2023.2252780. Gut Microbes. 2023. PMID: 37680117 Free PMC article. Review.
-
A master regulator of central carbon metabolism directly activates virulence gene expression in attaching and effacing pathogens.PLoS Pathog. 2024 Oct 15;20(10):e1012451. doi: 10.1371/journal.ppat.1012451. eCollection 2024 Oct. PLoS Pathog. 2024. PMID: 39405360 Free PMC article.
-
Quorum sensing in human gut and food microbiomes: Significance and potential for therapeutic targeting.Front Microbiol. 2022 Nov 25;13:1002185. doi: 10.3389/fmicb.2022.1002185. eCollection 2022. Front Microbiol. 2022. PMID: 36504831 Free PMC article. Review.
-
Enterobacterales Biofilm-Specific Genes and Antimicrobial and Anti-Inflammatory Biomarkers in the Blood of Patients with Ischemic Heart Disease.Diagnostics (Basel). 2024 Mar 5;14(5):546. doi: 10.3390/diagnostics14050546. Diagnostics (Basel). 2024. PMID: 38473018 Free PMC article.
References
-
- Knutton S, Baldwin T, Williams PH, McNeish AS. 1989. Actin accumulation at sites of bacterial adhesion to tissue culture cells: basis of a new diagnostic test for enteropathogenic and enterohemorrhagic Escherichia coli. Infect Immun 57:1290–1298. doi:10.1128/IAI.57.4.1290-1298.1989. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

