Clostridioides difficile exploits toxin-mediated inflammation to alter the host nutritional landscape and exclude competitors from the gut microbiota
- PMID: 33469019
- PMCID: PMC7815924
- DOI: 10.1038/s41467-020-20746-4
Clostridioides difficile exploits toxin-mediated inflammation to alter the host nutritional landscape and exclude competitors from the gut microbiota
Abstract
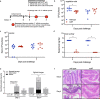
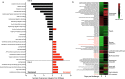
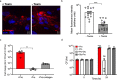
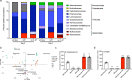
Clostridioides difficile is a bacterial pathogen that causes a range of clinical disease from mild to moderate diarrhea, pseudomembranous colitis, and toxic megacolon. Typically, C. difficile infections (CDIs) occur after antibiotic treatment, which alters the gut microbiota, decreasing colonization resistance against C. difficile. Disease is mediated by two large toxins and the expression of their genes is induced upon nutrient depletion via the alternative sigma factor TcdR. Here, we use tcdR mutants in two strains of C. difficile and omics to investigate how toxin-induced inflammation alters C. difficile metabolism, tissue gene expression and the gut microbiota, and to determine how inflammation by the host may be beneficial to C. difficile. We show that C. difficile metabolism is significantly different in the face of inflammation, with changes in many carbohydrate and amino acid uptake and utilization pathways. Host gene expression signatures suggest that degradation of collagen and other components of the extracellular matrix by matrix metalloproteinases is a major source of peptides and amino acids that supports C. difficile growth in vivo. Lastly, the inflammation induced by C. difficile toxin activity alters the gut microbiota, excluding members from the genus Bacteroides that are able to utilize the same essential nutrients released from collagen degradation.
Conflict of interest statement
C.M.T. consults for Vedanta Biosciences, Inc. and Summit Therapeutics.
Figures

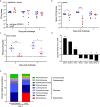
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

