Resonance Energy Transfer-Based Biosensors for Point-of-Need Diagnosis-Progress and Perspectives
- PMID: 33477883
- PMCID: PMC7833371
- DOI: 10.3390/s21020660
Resonance Energy Transfer-Based Biosensors for Point-of-Need Diagnosis-Progress and Perspectives
Abstract
The demand for point-of-need (PON) diagnostics for clinical and other applications is continuing to grow. Much of this demand is currently serviced by biosensors, which combine a bioanalytical sensing element with a transducing device that reports results to the user. Ideally, such devices are easy to use and do not require special skills of the end user. Application-dependent, PON devices may need to be capable of measuring low levels of analytes very rapidly, and it is often helpful if they are also portable. To date, only two transduction modalities, colorimetric lateral flow immunoassays (LFIs) and electrochemical assays, fully meet these requirements and have been widely adopted at the point-of-need. These modalities are either non-quantitative (LFIs) or highly analyte-specific (electrochemical glucose meters), therefore requiring considerable modification if they are to be co-opted for measuring other biomarkers. Förster Resonance Energy Transfer (RET)-based biosensors incorporate a quantitative and highly versatile transduction modality that has been extensively used in biomedical research laboratories. RET-biosensors have not yet been applied at the point-of-need despite its advantages over other established techniques. In this review, we explore and discuss recent developments in the translation of RET-biosensors for PON diagnoses, including their potential benefits and drawbacks.
Keywords: BRET; CRET; FRET; PADs; microfluidics; on-site; on-the-spot; point-of-care; time-resolved FRET.
Conflict of interest statement
K.C. and S.T. are listed as inventors in patents submitted by the CSIRO related to the Cybertongue® technology. S.T. is a founder of PPB Technology, which has licensed certain commercial rights to those patents.
Figures
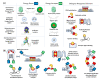




References
-
- Pontius K., Semenova D., Silina Y.E., Gernaey K.V., Junicke H. Automated Electrochemical Glucose Biosensor Platform as an Efficient Tool Toward On-Line Fermentation Monitoring: Novel Application Approaches and Insights. Front. Bioeng. Biotechnol. 2020;8:15. doi: 10.3389/fbioe.2020.00436. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

