The Landscape of Recombination Events That Create Nonribosomal Peptide Diversity
- PMID: 33480992
- PMCID: PMC8097286
- DOI: 10.1093/molbev/msab015
The Landscape of Recombination Events That Create Nonribosomal Peptide Diversity
Abstract
Nonribosomal peptides (NRP) are crucial molecular mediators in microbial ecology and provide indispensable drugs. Nevertheless, the evolution of the flexible biosynthetic machineries that correlates with the stunning structural diversity of NRPs is poorly understood. Here, we show that recombination is a key driver in the evolution of bacterial NRP synthetase (NRPS) genes across distant bacterial phyla, which has guided structural diversification in a plethora of NRP families by extensive mixing and matching of biosynthesis genes. The systematic dissection of a large number of individual recombination events did not only unveil a striking plurality in the nature and origin of the exchange units but allowed the deduction of overarching principles that enable the efficient exchange of adenylation (A) domain substrates while keeping the functionality of the dynamic multienzyme complexes. In the majority of cases, recombination events have targeted variable portions of the Acore domains, yet domain interfaces and the flexible Asub domain remained untapped. Our results strongly contradict the widespread assumption that adenylation and condensation (C) domains coevolve and significantly challenge the attributed role of C domains as stringent selectivity filter during NRP synthesis. Moreover, they teach valuable lessons on the choice of natural exchange units in the evolution of NRPS diversity, which may guide future engineering approaches.
Keywords: evolution; microbial ecology; natural products; nonribosomal peptide synthetases; recombination; structural diversity.
© The Author(s) 2021. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures





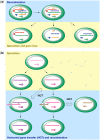
References
-
- Ackerley DF. 2016. Cracking the nonribosomal code. Cell Chem Biol. 23(5):535–537. - PubMed
-
- Alanjary M, Cano-Prieto C, Gross H, Medema MH.. 2019. Computer-aided re-engineering of nonribosomal peptide and polyketide biosynthetic assembly lines. Nat Prod Rep. 36(9):1249–1261. - PubMed
-
- Bachmann BO, Ravel J.. 2009. Chapter 8. Methods for in silico prediction of microbial polyketide and nonribosomal peptide biosynthetic pathways from DNA sequence data. Methods Enzymol. 458:181–217. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

