A disorder-related variant (E420K) of a PP2A-regulatory subunit (PPP2R5D) causes constitutively active AKT-mTOR signaling and uncoordinated cell growth
- PMID: 33482199
- PMCID: PMC7952134
- DOI: 10.1016/j.jbc.2021.100313
A disorder-related variant (E420K) of a PP2A-regulatory subunit (PPP2R5D) causes constitutively active AKT-mTOR signaling and uncoordinated cell growth
Abstract
Functional genomic approaches have facilitated the discovery of rare genetic disorders and improved efforts to decipher their underlying etiology. PPP2R5D-related disorder is an early childhood onset condition characterized by intellectual disability, hypotonia, autism-spectrum disorder, macrocephaly, and dysmorphic features. The disorder is caused by de novo single nucleotide changes in PPP2R5D, which generate heterozygous dominant missense variants. PPP2R5D is known to encode a B'-type (B'56δ) regulatory subunit of a PP2A-serine/threonine phosphatase. To help elucidate the molecular mechanisms altered in PPP2R5D-related disorder, we used a CRISPR-single-base editor to generate HEK-293 cells in which a single transition (c.1258G>A) was introduced into one allele, precisely recapitulating a clinically relevant E420K variant. Unbiased quantitative proteomic and phosphoproteomic analyses of endogenously expressed proteins revealed heterozygous-dominant changes in kinase/phosphatase signaling. These data combined with orthogonal validation studies revealed a previously unrecognized interaction of PPP2R5D with AKT in human cells, leading to constitutively active AKT-mTOR signaling, increased cell size, and uncoordinated cellular growth in E420K-variant cells. Rapamycin reduced cell size and dose-dependently reduced RPS6 phosphorylation in E420K-variant cells, suggesting that inhibition of mTOR1 can suppress both the observed RPS6 hyperphosphorylation and increased cell size. Together, our findings provide a deeper understanding of PPP2R5D and insight into how the E420K-variant alters signaling networks influenced by PPP2R5D. Our comprehensive approach, which combines precise genome editing, isobaric tandem mass tag labeling of peptides generated from endogenously expressed proteins, and concurrent liquid chromatography-mass spectrometry (LC-MS3), also provides a roadmap that can be used to rapidly explore the etiologies of additional genetic disorders.
Keywords: AKT-mTOR; Jordan’s syndrome; PP2A; PPP2R5D; PPP2R5D-intellectual disability; PPP2R5D-related neurodevelopmental disorder; phosphatase.
Copyright © 2021 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflicts of interest with the contents of this article.
Figures

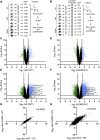





References
-
- Houge G., Haesen D., Vissers L.E., Mehta S., Parker M.J., Wright M., Vogt J., McKee S., Tolmie J.L., Cordeiro N., Kleefstra T., Willemsen M.H., Reijnders M.R., Berland S., Hayman E. B56delta-related protein phosphatase 2A dysfunction identified in patients with intellectual disability. J. Clin. Invest. 2015;125:3051–3062. - PMC - PubMed
-
- Shang L., Henderson L.B., Cho M.T., Petrey D.S., Fong C.T., Haude K.M., Shur N., Lundberg J., Hauser N., Carmichael J., Innis J., Schuette J., Wu Y.W., Asaikar S., Pearson M. De novo missense variants in PPP2R5D are associated with intellectual disability, macrocephaly, hypotonia, and autism. Neurogenetics. 2016;17:43–49. - PMC - PubMed
-
- Loveday C., Tatton-Brown K., Clarke M., Westwood I., Renwick A., Ramsay E., Nemeth A., Campbell J., Joss S., Gardner M., Zachariou A., Elliott A., Ruark E., van Montfort R., Childhood Overgrowth C. Mutations in the PP2A regulatory subunit B family genes PPP2R5B, PPP2R5C and PPP2R5D cause human overgrowth. Hum. Mol. Genet. 2015;24:4775–4779. - PMC - PubMed
-
- Mirzaa G., Foss K., Nattakom M., Chung W.K. PPP2R5D-Related neurodevelopmental disorder. In: Adam M.P., Ardinger H.H., Pagon R.A., Wallace S.E., Bean L.J.H., Stephens K., Amemiya A., editors. GeneReviews [Internet] University of Washington, Seattle; Seattle, WA: 2019. pp. 1193–2020.
-
- Deciphering Developmental Disorders Study. Fitzgerald T.W., Gerety S.S., Jones W.D., van Kogelenberg M., King D.A., McRae J., Morley K.I., Parthiban V., Al-Turki S., Ambridge K., Barrett D.M., Bayzetinova T., Clayton S., Coomber E.L., Gribble S. Large-scale discovery of novel genetic causes of developmental disorders. Nature. 2015;519:223–228. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

