Spanning the gap: unraveling RSC dynamics in vivo
- PMID: 33484328
- PMCID: PMC8139908
- DOI: 10.1007/s00294-020-01144-1
Spanning the gap: unraveling RSC dynamics in vivo
Abstract
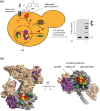
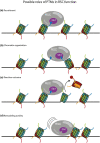
Multiple reports over the past 2 years have provided the first complete structural analyses for the essential yeast chromatin remodeler, RSC, providing elaborate molecular details for its engagement with the nucleosome. However, there still remain gaps in resolution, particularly within the many RSC subunits that harbor histone binding domains.Solving contacts at these interfaces is crucial because they are regulated by posttranslational modifications that control remodeler binding modes and function. Modifications are dynamic in nature often corresponding to transcriptional activation states and cell cycle stage, highlighting not only a need for enriched spatial resolution but also temporal understanding of remodeler engagement with the nucleosome. Our recent work sheds light on some of those gaps by exploring the binding interface between the RSC catalytic motor protein, Sth1, and the nucleosome, in the living nucleus. Using genetically encoded photo-activatable amino acids incorporated into histones of living yeast we are able to monitor the nucleosomal binding of RSC, emphasizing the regulatory roles of histone modifications in a spatiotemporal manner. We observe that RSC prefers to bind H2B SUMOylated nucleosomes in vivo and interacts with neighboring nucleosomes via H3K14ac. Additionally, we establish that RSC is constitutively bound to the nucleosome and is not ejected during mitotic chromatin compaction but alters its binding mode as it progresses through the cell cycle. Our data offer a renewed perspective on RSC mechanics under true physiological conditions.
Keywords: Chromatin remodelling; Genetic code expansion; Lysine acetylation; Photo-crosslinking; RSC; Sumoylation; Unnatural amino acids.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
Similar articles
-
Architecture of the chromatin remodeler RSC and insights into its nucleosome engagement.Elife. 2019 Dec 30;8:e54449. doi: 10.7554/eLife.54449. Elife. 2019. PMID: 31886770 Free PMC article.
-
Regulation of DNA Translocation Efficiency within the Chromatin Remodeler RSC/Sth1 Potentiates Nucleosome Sliding and Ejection.Mol Cell. 2016 May 5;62(3):453-461. doi: 10.1016/j.molcel.2016.03.032. Mol Cell. 2016. PMID: 27153540 Free PMC article.
-
The chromatin remodelers RSC and ISW1 display functional and chromatin-based promoter antagonism.Elife. 2015 Mar 30;4:e06073. doi: 10.7554/eLife.06073. Elife. 2015. PMID: 25821983 Free PMC article.
-
Establishing nucleosome architecture and stability at promoters: Roles of pioneer transcription factors and the RSC chromatin remodeler.Bioessays. 2017 May;39(5). doi: 10.1002/bies.201600237. Epub 2017 Mar 27. Bioessays. 2017. PMID: 28345796 Review.
-
A conserved role of the RSC chromatin remodeler in the establishment of nucleosome-depleted regions.Curr Genet. 2017 May;63(2):187-193. doi: 10.1007/s00294-016-0642-y. Epub 2016 Aug 24. Curr Genet. 2017. PMID: 27558480 Free PMC article. Review.
Cited by
-
Unnatural Amino Acid Crosslinking for Increased Spatiotemporal Resolution of Chromatin Dynamics.Int J Mol Sci. 2023 Aug 17;24(16):12879. doi: 10.3390/ijms241612879. Int J Mol Sci. 2023. PMID: 37629060 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

