The nonhuman primate neuroimaging and neuroanatomy project
- PMID: 33484849
- PMCID: PMC8079967
- DOI: 10.1016/j.neuroimage.2021.117726
The nonhuman primate neuroimaging and neuroanatomy project
Abstract
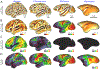
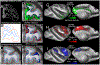
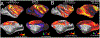
Multi-modal neuroimaging projects such as the Human Connectome Project (HCP) and UK Biobank are advancing our understanding of human brain architecture, function, connectivity, and their variability across individuals using high-quality non-invasive data from many subjects. Such efforts depend upon the accuracy of non-invasive brain imaging measures. However, 'ground truth' validation of connectivity using invasive tracers is not feasible in humans. Studies using nonhuman primates (NHPs) enable comparisons between invasive and non-invasive measures, including exploration of how "functional connectivity" from fMRI and "tractographic connectivity" from diffusion MRI compare with long-distance connections measured using tract tracing. Our NonHuman Primate Neuroimaging & Neuroanatomy Project (NHP_NNP) is an international effort (6 laboratories in 5 countries) to: (i) acquire and analyze high-quality multi-modal brain imaging data of macaque and marmoset monkeys using protocols and methods adapted from the HCP; (ii) acquire quantitative invasive tract-tracing data for cortical and subcortical projections to cortical areas; and (iii) map the distributions of different brain cell types with immunocytochemical stains to better define brain areal boundaries. We are acquiring high-resolution structural, functional, and diffusion MRI data together with behavioral measures from over 100 individual macaques and marmosets in order to generate non-invasive measures of brain architecture such as myelin and cortical thickness maps, as well as functional and diffusion tractography-based connectomes. We are using classical and next-generation anatomical tracers to generate quantitative connectivity maps based on brain-wide counting of labeled cortical and subcortical neurons, providing ground truth measures of connectivity. Advanced statistical modeling techniques address the consistency of both kinds of data across individuals, allowing comparison of tracer-based and non-invasive MRI-based connectivity measures. We aim to develop improved cortical and subcortical areal atlases by combining histological and imaging methods. Finally, we are collecting genetic and sociality-associated behavioral data in all animals in an effort to understand how genetic variation shapes the connectome and behavior.
Keywords: Connectivity; Connectome; Diffusion MRI; Functional MRI; Hierarchy; Human; Macaque; Marmoset; Retrograde tracer.
Copyright © 2021 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Competing Interest All the authors have no conflicts of interest to declare.
Figures


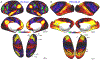


References
-
- Autio JA, Glasser MF, Ose T, Donahue CJ, Bastiani M, Ohno M, Kawabata Y, Urushibata Y, Murata K, Nishigori K, Yamaguchi M, Hori Y, Yoshida A, Go Y, Coalson TS, Jbabdi S, Sotiropoulos SN, Kennedy H, Smith S, Van Essen DC, Hayashi T, 2020b. Towards HCP-Style Macaque Connectomes: 24-Channel 3T Multi-Array Coil, MRI Sequences and Preprocessing. NeuroImage, 116800 doi: 10.1016/j.neuroimage.2020.116800. - DOI - PMC - PubMed
-
- Bakken TE, Miller JA, Ding S-L, Sunkin SM, Smith KA, Ng L, Szafer A, Dalley RA, Royall JJ, Lemon T, Shapouri S, Aiona K, Arnold J, Bennett JL, Bertagnolli D, Bickley K, Boe A, Brouner K, Butler S, Byrnes E, Caldejon S, Carey A, Cate S, Chapin M, Chen J, Dee N, Desta T, Dolbeare TA, Dotson N, Ebbert A, Fulfs E, Gee G, Gilbert TL, Goldy J, Gourley L, Gregor B, Gu G, Hall J, Haradon Z, Haynor DR, Hejazinia N, Hoerder-Suabedissen A, Howard R, Jochim J, Kinnunen M, Kriedberg A, Kuan CL, Lau C, Lee C-K, Lee F, Luong L, Mastan N, May R, Melchor J, Mosqueda N, Mott E, Ngo K, Nyhus J, Oldre A, Olson E, Parente J, Parker PD, Parry S, Pendergraft J, Potekhina L, Reding M, Riley ZL, Roberts T, Rogers B, Roll K, Rosen D, Sandman D, Sarreal M, Shapovalova N, Shi S, Sjoquist N, Sodt AJ, Townsend R, Velasquez L, Wagley U, Wakeman WB, White C, Bennett C, Wu J, Young R, Youngstrom BL, Wohnoutka P, Gibbs RA, Rogers J, Hohmann JG, Hawrylycz MJ, Hevner RF, Molnár Z, Phillips JW, Dang C, Jones AR, Amaral DG, Bernard A, Lein ES, 2016. A comprehensive transcriptional map of primate brain development. Nature 535, 367–375. doi: 10.1038/nature18637. - DOI - PMC - PubMed
References under revision of this special issue of NeuroImage
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

