FOS licenses early events in stem cell activation driving skeletal muscle regeneration
- PMID: 33503437
- PMCID: PMC9112118
- DOI: 10.1016/j.celrep.2020.108656
FOS licenses early events in stem cell activation driving skeletal muscle regeneration
Abstract
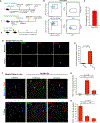
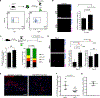
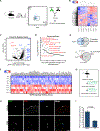
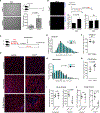
Muscle satellite cells (SCs) are a quiescent (non-proliferative) stem cell population in uninjured skeletal muscle. Although SCs have been investigated for nearly 60 years, the molecular drivers that transform quiescent SCs into the rapidly dividing (activated) stem/progenitor cells that mediate muscle repair after injury remain largely unknown. Here we identify a prominent FBJ osteosarcoma oncogene (Fos) mRNA and protein signature in recently activated SCs that is rapidly, heterogeneously, and transiently induced by muscle damage. We further reveal a requirement for FOS to efficiently initiate key stem cell functions, including cell cycle entry, proliferative expansion, and muscle regeneration, via induction of "pro-regenerative" target genes that stimulate cell migration, division, and differentiation. Disruption of one of these Fos/AP-1 targets, NAD(+)-consuming mono-ADP-ribosyl-transferase 1 (Art1), in SCs delays cell cycle entry and impedes progenitor cell expansion and muscle regeneration. This work uncovers an early-activated FOS/ART1/mono-ADP-ribosylation (MARylation) pathway that is essential for stem cell-regenerative responses.
Keywords: AP-1; ART1; FOS; MARylation; early activation; muscle satellite cells; muscle stem cells; post-translational regulation.
Copyright © 2021 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests A.J.W. is a scientific advisor for Frequency Therapeutics, and A.J.W. and L.L.R. are co-founders and scientific advisory board members and hold private equity in Elevian, Inc., a company that aims to develop medicines to restore regenerative capacity. Elevian also provides sponsored research to the Wagers lab.
Figures


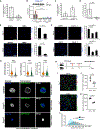
References
-
- Bolstad BM, Irizarry RA, Astrand M, and Speed TP (2003). A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

