Identification and molecular characterization of mutations in nucleocapsid phosphoprotein of SARS-CoV-2
- PMID: 33505806
- PMCID: PMC7789862
- DOI: 10.7717/peerj.10666
Identification and molecular characterization of mutations in nucleocapsid phosphoprotein of SARS-CoV-2
Abstract
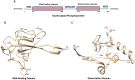
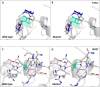
SARS-CoV-2 genome encodes four structural proteins that include the spike glycoprotein, membrane protein, envelope protein and nucleocapsid phosphoprotein (N-protein). The N-protein interacts with viral genomic RNA and helps in packaging. As SARS-CoV-2 spread to almost all countries worldwide within 2-3 months, it also acquired mutations in its RNA genome. Therefore, this study was conducted with an aim to identify the variations present in N-protein of SARS-CoV-2. Here, we analysed 4,163 reported sequence of N-protein from United States of America (USA) and compared them with the first reported sequence from Wuhan, China. Our study identified 107 mutations that reside all over the N-protein. Further, we show the high rate of mutations in intrinsically disordered regions (IDRs) of N-protein. Our study show 45% residues of IDR2 harbour mutations. The RNA-binding domain (RBD) and dimerization domain of N-protein also have mutations at key residues. We further measured the effect of these mutations on N-protein stability and dynamicity and our data reveals that multiple mutations can cause considerable alterations. Altogether, our data strongly suggests that N-protein is one of the mutational hotspot proteins of SARS-CoV-2 that is changing rapidly and these mutations can potentially interferes with various aspects of N-protein functions including its interaction with RNA, oligomerization and signalling events.
Keywords: COVID-19; Infectious disease; Mutations; Nucleocapsid Phosphoprotein (N protein); SARS-CoV-2; USA.
© 2021 Azad.
Conflict of interest statement
The author declares that they have no competing interests.
Figures


References
-
- Blanco-Melo D, Nilsson-Payant BE, Liu WC, Uhl S, Hoagland D, Møller R, Jordan TX, Oishi K, Panis M, Sachs D, Wang TT, Schwartz RE, Lim JK, Albrecht RA, TenOever BR. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell. 2020;181(5):1036–1045.e9. doi: 10.1016/j.cell.2020.04.026. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

