Evaluation of a multiplex PCR method for the detection of porcine parvovirus types 1 through 7 using various field samples
- PMID: 33508002
- PMCID: PMC7842984
- DOI: 10.1371/journal.pone.0245699
Evaluation of a multiplex PCR method for the detection of porcine parvovirus types 1 through 7 using various field samples
Abstract
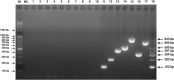
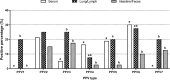
Porcine parvoviruses (PPVs) are small, nonenveloped DNA viruses that are widespread in the global pig population. PPV type 1 (PPV1) is a major causative agent of reproductive failure and has been recognized since the 1960s. In recent decades, novel PPVs have been identified and designated as PPVs 2 through 7 (PPV2~PPV7). Although the epidemiological impacts of these newly recognized parvoviruses on pigs are largely unknown, continuous surveillance of these PPVs is needed. The aim of this study was to develop an improved and efficient detection tool for these PPVs and to assess the developed method with field samples. Using 7 sets of newly designed primers, a multiplex polymerase chain reaction (mPCR) protocol was developed for the simultaneous detection of the seven genotypes of PPV (PPV1~PPV7). The sensitivity of the mPCR assay was analyzed, and the detection limit was determined to be 3×103 viral copies. The assay was highly specific in detecting one or more of the viruses in various combinations in specimens. The mPCR method was evaluated with 80 serum samples, 40 lung or lymph node samples and 40 intestine or fecal samples. When applied to these samples, the mPCR method could detect the 7 viruses simultaneously, providing rapid results regarding infection and coinfection status. In conclusion, the developed mPCR assay can be utilized as an effective and accurate diagnostic tool for rapid differential detection and epidemiological surveillance of various PPVs in numerous types of field samples.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
A Systematic Investigation Unveils High Coinfection Status of Porcine Parvovirus Types 1 through 7 in China from 2016 to 2020.Microbiol Spectr. 2021 Dec 22;9(3):e0129421. doi: 10.1128/Spectrum.01294-21. Epub 2021 Dec 1. Microbiol Spectr. 2021. PMID: 34851175 Free PMC article.
-
Detection Patterns of Porcine Parvovirus (PPV) and Novel Porcine Parvoviruses 2 through 6 (PPV2-PPV6) in Polish Swine Farms.Viruses. 2019 May 24;11(5):474. doi: 10.3390/v11050474. Viruses. 2019. PMID: 31137628 Free PMC article.
-
Do porcine parvoviruses 1 through 7 (PPV1-PPV7) have an impact on porcine circovirus type 2 (PCV2) viremia in pigs?Vet Microbiol. 2020 Mar;242:108613. doi: 10.1016/j.vetmic.2020.108613. Epub 2020 Feb 17. Vet Microbiol. 2020. PMID: 32122579
-
The Novel Porcine Parvoviruses: Current State of Knowledge and Their Possible Implications in Clinical Syndromes in Pigs.Viruses. 2023 Dec 9;15(12):2398. doi: 10.3390/v15122398. Viruses. 2023. PMID: 38140639 Free PMC article. Review.
-
Molecular epidemiology and evolution of porcine parvoviruses.Infect Genet Evol. 2015 Dec;36:300-306. doi: 10.1016/j.meegid.2015.10.007. Epub 2015 Oct 22. Infect Genet Evol. 2015. PMID: 26453771 Review.
Cited by
-
Infectious Agents Associated with Abortion Outbreaks in Italian Pig Farms from 2011 to 2021.Vet Sci. 2024 Oct 12;11(10):496. doi: 10.3390/vetsci11100496. Vet Sci. 2024. PMID: 39453088 Free PMC article.
-
Emergent Molecular Techniques Applied to the Detection of Porcine Viruses.Vet Sci. 2023 Oct 7;10(10):609. doi: 10.3390/vetsci10100609. Vet Sci. 2023. PMID: 37888561 Free PMC article. Review.
-
Prevalence of porcine parvovirus 1 through 7 (PPV1-PPV7) and co-factor association with PCV2 and PRRSV in Korea.BMC Vet Res. 2022 Apr 9;18(1):133. doi: 10.1186/s12917-022-03236-1. BMC Vet Res. 2022. PMID: 35395853 Free PMC article.
-
Identification and in vitro characterization of a novel porcine parvovirus 6 in Russia.Vet Res Commun. 2024 Feb;48(1):417-425. doi: 10.1007/s11259-023-10226-7. Epub 2023 Sep 29. Vet Res Commun. 2024. PMID: 37773486
-
First Molecular Characterisation of Porcine Parvovirus 7 (PPV7) in Italy.Viruses. 2024 Jun 8;16(6):932. doi: 10.3390/v16060932. Viruses. 2024. PMID: 38932224 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

