Therapeutic shutdown of HBV transcripts promotes reappearance of the SMC5/6 complex and silencing of the viral genome in vivo
- PMID: 33509930
- PMCID: PMC8762019
- DOI: 10.1136/gutjnl-2020-322571
Therapeutic shutdown of HBV transcripts promotes reappearance of the SMC5/6 complex and silencing of the viral genome in vivo
Abstract
Objective: Therapeutic strategies silencing and reducing the hepatitis B virus (HBV) reservoir, the covalently closed circular DNA (cccDNA), have the potential to cure chronic HBV infection. We aimed to investigate the impact of small interferring RNA (siRNA) targeting all HBV transcripts or pegylated interferon-α (peg-IFNα) on the viral regulatory HBx protein and the structural maintenance of chromosome 5/6 complex (SMC5/6), a host factor suppressing cccDNA transcription. In particular, we assessed whether interventions lowering HBV transcripts can achieve and maintain silencing of cccDNA transcription in vivo.
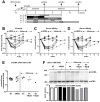
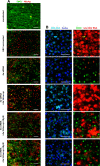
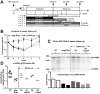
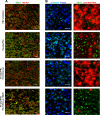
Design: HBV-infected human liver chimeric mice were treated with siRNA or peg-IFNα. Virological and host changes were analysed at the end of treatment and during the rebound phase by qualitative PCR, ELISA, immunoblotting and chromatin immunoprecipitation. RNA in situ hybridisation was combined with immunofluorescence to detect SMC6 and HBV RNAs at single cell level. The entry inhibitor myrcludex-B was used during the rebound phase to avoid new infection events.
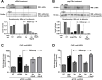
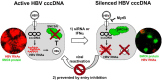
Results: Both siRNA and peg-IFNα strongly reduced all HBV markers, including HBx levels, thus enabling the reappearance of SMC5/6 in hepatocytes that achieved HBV-RNA negativisation and SMC5/6 association with the cccDNA. Only IFN reduced cccDNA loads and enhanced IFN-stimulated genes. However, the antiviral effects did not persist off treatment and SMC5/6 was again degraded. Remarkably, the blockade of viral entry that started at the end of treatment hindered renewed degradation of SMC5/6.
Conclusion: These results reveal that therapeutics abrogating all HBV transcripts including HBx promote epigenetic suppression of the HBV minichromosome, whereas strategies protecting the human hepatocytes from reinfection are needed to maintain cccDNA silencing.
Keywords: antiviral therapy; hepatitis B; interferon-α; liver.
© Author(s) (or their employer(s)) 2022. Re-use permitted under CC BY-NC. No commercial re-use. See rights and permissions. Published by BMJ.
Conflict of interest statement
Competing interests: RCM, RKB, HJ and SPF are employees of Gilead Sciences. SU is co-applicant and co-inventor on patents protecting myrcludex-B (bulevirtide). The other authors have nothing to disclose.
Figures
Comment in
-
Silencing HBV transcription with SMC5/6: has a path been found?Gut. 2022 Feb;71(2):233-234. doi: 10.1136/gutjnl-2021-324144. Epub 2021 Mar 1. Gut. 2022. PMID: 33649046 No abstract available.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
