The Ustilago hordei-Barley Interaction Is a Versatile System for Characterization of Fungal Effectors
- PMID: 33513785
- PMCID: PMC7912019
- DOI: 10.3390/jof7020086
The Ustilago hordei-Barley Interaction Is a Versatile System for Characterization of Fungal Effectors
Abstract
Obligate biotrophic fungal pathogens, such as Blumeria graminis and Puccinia graminis, are amongst the most devastating plant pathogens, causing dramatic yield losses in many economically important crops worldwide. However, a lack of reliable tools for the efficient genetic transformation has hampered studies into the molecular basis of their virulence or pathogenicity. In this study, we present the Ustilago hordei-barley pathosystem as a model to characterize effectors from different plant pathogenic fungi. We generate U. hordei solopathogenic strains, which form infectious filaments without the presence of a compatible mating partner. Solopathogenic strains are suitable for heterologous expression system for fungal virulence factors. A highly efficient Crispr/Cas9 gene editing system is made available for U. hordei. In addition, U. hordei infection structures during barley colonization are analyzed using transmission electron microscopy, showing that U. hordei forms intracellular infection structures sharing high similarity to haustoria formed by obligate rust and powdery mildew fungi. Thus, U. hordei has high potential as a fungal expression platform for functional studies of heterologous effector proteins in barley.
Keywords: CRISPR-Cas9; Ustilago hordei; effectors; haustoria; heterologous gene expression.
Conflict of interest statement
The authors have no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures



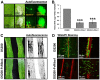
References
-
- Spanu P.D., Abbott J.C., Amselem J., Burgis T.A., Soanes D.M., Stüber K., Van Themaat E.V.L., Brown J.K.M., Butcher S.A., Gurr S.J., et al. Genome expansion and gene loss in powdery mildew fungi reveal tradeoffs in extreme parasitism. Science. 2010;330:1543–1546. doi: 10.1126/science.1194573. - DOI - PubMed
-
- Hacquard S., Joly D.L., Lin Y.-C., Tisserant E., Feau N., Delaruelle C., Legué V., Kohler A., Tanguay P., Petre B., et al. A Comprehensive analysis of genes encoding small secreted proteins identifies candidate effectors in Melampsora larici-populina (Poplar Leaf Rust) Mol. Plant Microbe Interact. 2012;25:279–293. doi: 10.1094/MPMI-09-11-0238. - DOI - PubMed
-
- Duplessis S., Cuomo C.A., Lin Y.-C., Aerts A., Tisserant E., Veneault-Fourrey C., Joly D.L., Hacquard S., Amselem J., Cantarel B.L., et al. Obligate biotrophy features unraveled by the genomic analysis of rust fungi. Proc. Natl. Acad. Sci. USA. 2011;108:9166–9171. doi: 10.1073/pnas.1019315108. - DOI - PMC - PubMed
-
- O’Connell R.J., Thon M.R., Hacquard S., Amyotte S.G., Kleemann J., Torres M.F., Damm U., Buiate E.A., Epstein L., Alkan N., et al. Lifestyle transitions in plant pathogenic Colletotrichum fungi deciphered by genome and transcriptome analyses. Nat. Genet. 2012;44:1060–1065. doi: 10.1038/ng.2372. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

